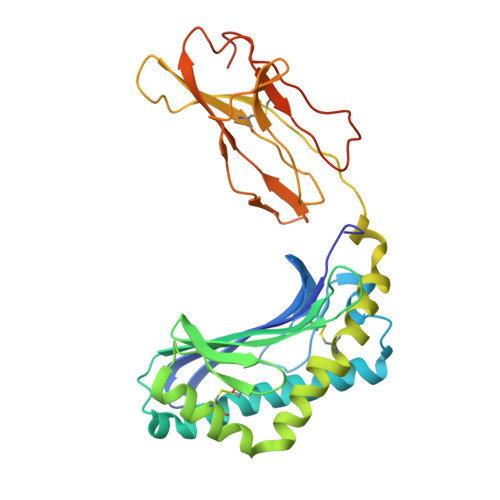
The crystal structure of human CD1b with a bound bacterial glycolipid.
Batuwangala, T., Shepherd, D., Gadola, S.D., Gibson, K.J., Zaccai, N.R., Fersht, A.R., Besra, G.S., Cerundolo, V., Jones, E.Y.(2004) J Immunol 172: 2382-2388
- PubMed: 14764708 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.172.4.2382
- Primary Citation Related Structures:
1UQS - PubMed Abstract:
The human MHC class I-like molecule CD1b is distinctive among CD1 alleles in that it is capable of presenting a set of glycolipid species that show a very broad range of variation in the lengths of their acyl chains. A structure of CD1b complexed with relatively short acyl chain glycolipids plus detergent suggested how an interlinked network of channels within the Ag-binding groove could accommodate acyl chain lengths of up to 80 carbons. The structure of CD1b complexed with glucose monomycolate, reported in this study, confirms this hypothesis and illustrates how the distinctive substituents of intracellular bacterial glycolipids can be accommodated. The Ag-binding groove of CD1b is, uniquely among CD1 alleles, partitioned into channels suitable for the compact accommodation of lengthy acyl chains. The current crystal structure illustrates for the first time the binding of a natural bacterial lipid Ag to CD1b and shows how its novel structural features fit this molecule for its role in the immune response to intracellular bacteria.
- Cancer Research UK Receptor Structure Group, The Division of Structural Biology, and Cancer Research UK.
Organizational Affiliation: