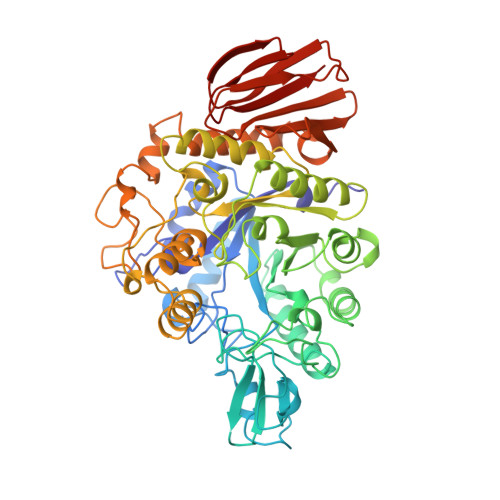
The refined crystal structure of Bacillus cereus oligo-1,6-glucosidase at 2.0 A resolution: structural characterization of proline-substitution sites for protein thermostabilization.
Watanabe, K., Hata, Y., Kizaki, H., Katsube, Y., Suzuki, Y.(1997) J Mol Biology 269: 142-153
- PubMed: 9193006
- DOI: https://doi.org/10.1006/jmbi.1997.1018
- Primary Citation Related Structures:
1UOK - PubMed Abstract:
The crystal structure of oligo-1,6-glucosidase (dextrin 6-alpha-glucanohydrolase, EC 3.2.1.10) from Bacillus cereus ATCC7064 has been refined to 2.0 A resolution with an R-factor of 19.6% for 43,328 reflections. The final model contains 4646 protein atoms and 221 ordered water molecules with respective root-mean-square deviations of 0.015 A for bond lengths and of 3.166 degrees for bond angles from the ideal values. The structure consists of three domains: the N-terminal domain (residues 1 to 104 and 175 to 480), the subdomain (residues 105 to 174) and the C-terminal domain (residues 481 to 558). The N-terminal domain takes a (beta/alpha)8-barrel structure with additions of an alpha-helix (N alpha6') between the sixth strand Nbeta6 and the sixth helix N alpha6, an alpha-helix (N alpha7') between the seventh strand Nbeta7 and the seventh helix N alpha7 and three alpha-helices (N alpha8', N alpha8" and N alpha8'" between the eighth strand Nbeta8 and the eighth helix N alpha8. The subdomain is composed of an alpha-helix, a three-stranded antiparallel beta-sheet, and long intervening loops. The C-terminal domain has a beta-barrel structure of eight antiparallel beta-strands folded in double Greek key motifs, which is distorted in the sixth strand Cbeta6. Three catalytic residues, Asp199, Glu255 and Asp329, are located at the bottom of a deep cleft formed by the subdomain and a cluster of the two additional alpha-helices N alpha8' and N alpha8" in the (beta/alpha)8-barrel. The refined structure reveals the locations of 21 proline-substitution sites that are expected to be critical to protein thermostabilization from a sequence comparison among three Bacillus oligo-1,6-glucosidases with different thermostability. These sites lie in loops, beta-turns and alpha-helices, in order of frequency, except for Cys515 in the fourth beta-strand Cbeta4 of the C-terminal domain. The residues in beta-turns (Lys121, Glu208, Pro257, Glu290, Pro443, Lys457 and Glu487) are all found at their second positions, and those in alpha-helices (Asn109, Glu175, Thr261 and Ile403) are present at their N1 positions of the first helical turns. Those residues in both secondary structures adopt phi and phi values favorable for proline substitution. Residues preceding the 21 sites are mostly conserved upon proline occurrence at these 21 sites in more thermostable Bacillus oligo-1,6-glucosidases. These structural features with respect to the 21 sites indicate that the sites in beta-turns and alpha-helices have more essential prerequisites for proline substitution to thermostabilize the protein than those in loops. This well supports the previous finding that the replacement at the appropriate positions in beta-turns or alpha-helices is the most effective for protein thermostabilization by proline substitution.
- Department of Agricultural Chemistry, Kyoto Prefectural University, Shimogamo, Sakyo, Japan.
Organizational Affiliation: