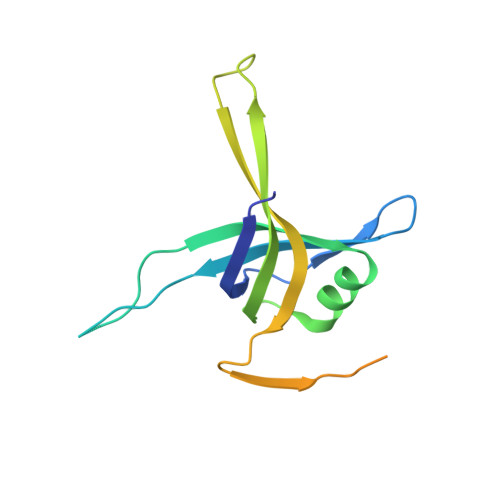
Structure of Mycobacterium tuberculosis single-stranded DNA-binding protein. Variability in quaternary structure and its implications
Saikrishnan, K., Jeyakanthan, J., Venkatesh, J., Acharya, N., Sekar, K., Varshney, U., Vijayan, M.(2003) J Mol Biology 331: 385-393
- PubMed: 12888346 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00729-0
- Primary Citation Related Structures:
1UE1, 1UE5, 1UE6, 1UE7 - PubMed Abstract:
Single-stranded DNA-binding protein (SSB) is an essential protein necessary for the functioning of the DNA replication, repair and recombination machineries. Here we report the structure of the DNA-binding domain of Mycobacterium tuberculosis SSB (MtuSSB) in four different crystals distributed in two forms. The structure of one of the forms was solved by a combination of isomorphous replacement and anomalous scattering. This structure was used to determine the structure of the other form by molecular replacement. The polypeptide chain in the structure exhibits the oligonucleotide binding fold. The globular core of the molecule in different subunits in the two forms and those in Escherichia coli SSB (EcoSSB) and human mitochondrial SSB (HMtSSB) have similar structure, although the three loops exhibit considerable structural variation. However, the tetrameric MtuSSB has an as yet unobserved quaternary association. This quaternary structure with a unique dimeric interface lends the oligomeric protein greater stability, which may be of significance to the functioning of the protein under conditions of stress. Also, as a result of the variation in the quaternary structure the path adopted by the DNA to wrap around MtuSSB is expected to be different from that of EcoSSB.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560012, India.
Organizational Affiliation: