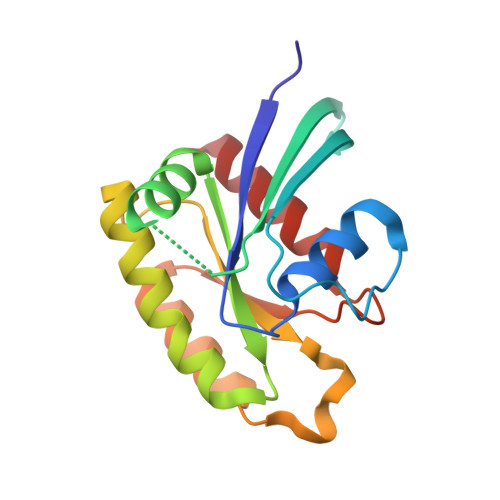
Crystal Structures of Ral-GppNHp and Ral-GDP Reveal Two Binding Sites that Are Also Present in Ras and Rap
Nicely, N.I., Kosak, J., de Serrano, V., Mattos, C.(2004) Structure 12: 2025-2036
- PubMed: 15530367 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.08.011
- Primary Citation Related Structures:
1U8Y, 1U8Z, 1U90 - PubMed Abstract:
RalA is a GTPase with effectors such as Sec5 and Exo84 in the exocyst complex and RalBP1, a GAP for Rho proteins. We report the crystal structures of Ral-GppNHp and Ral-GDP. Disordered switch I and switch II, located away from crystal contacts, are observed in one of the molecules in the asymmetric unit of the Ral-GppNHp structure. In the other molecule in the asymmetric unit, a second Mg(2+) ion is bound to the GppNHp gamma-phosphate in an environment in which switch I is pulled away from the nucleotide and switch II is found in a tight beta turn. Clustering of conserved residues on the surface of Ral-GppNHp identifies two putative sites for protein-protein interaction. One site is adjacent to switch I. The other is modulated by switch II and is obstructed in Ral-GDP. The Ral structures are discussed in the context of the published structures of the Ral/Sec5 complex, Ras, and Rap.
- Department of Molecular and Structural Biochemistry, 128 Polk Hall-CB 7622, North Carolina State University, Raleigh, NC 27695, USA.
Organizational Affiliation: