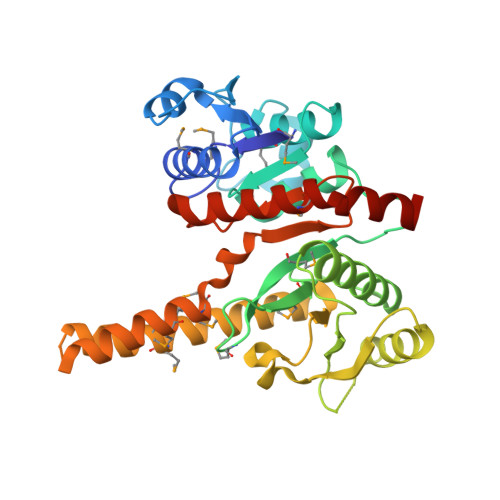
Crystal structure of fatty acid/phospholipid synthesis protein PlsX from Enterococcus faecalis.
Kim, Y., Li, H., Binkowski, T.A., Holzle, D., Joachimiak, A.(2009) J Struct Funct Genomics 10: 157-163
- PubMed: 19058030
- DOI: https://doi.org/10.1007/s10969-008-9052-9
- Primary Citation of Related Structures:
1U7N - PubMed Abstract:
PlsX is a key enzyme that coordinates the production of fatty acids and membrane phospholipids. The plsX gene is co-localized with a bacterial fab gene cluster which encodes several key fatty acid biosynthetic enzymes. The protein is a member of a large, conserved protein family (Pfam02504) found exclusively in bacteria. The PlsX sequence homologues include both phosphate acetyltransferases and phosphate butaryltransferases that catalyze the transfer of an acetyl or butaryl group to orthophosphate. We have determined the crystal structure of PlsX from the human pathogen Enterococcus faecalis. PlsX is a alpha/beta/alpha sandwich that resembles a Rossmann fold and forms a dimer. A putative catalytic site has been identified within a deep groove on the interface between monomers. This site showed strong surface similarity to epimerases and reductases. It was recently proposed that PlsX is a phosphate acyltransferase that catalyzes the formation of acyl-phosphate from the acyl-acyl carrier protein; however the specific biochemical function of the PlsX protein awaits further experimental scrutiny.
- Midwest Center for Structural Genomics and Structural Biology Center, Biosciences, Argonne National Laboratory, Argonne, IL 60439, USA.
Organizational Affiliation: