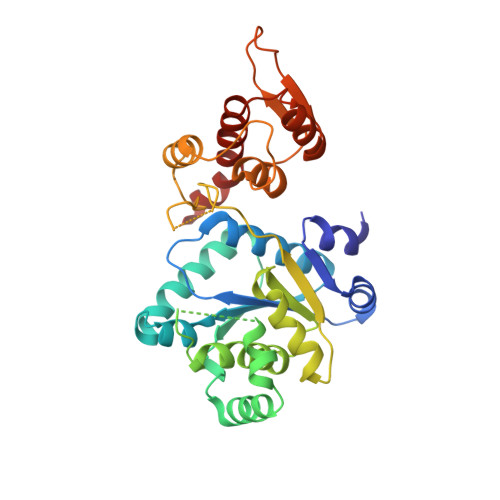
Crystal structures of apo wild-type M. jannaschii tyrosyl-tRNA synthetase (TyrRS) and an engineered TyrRS specific for O-methyl-L-tyrosine.
Zhang, Y., Wang, L., Schultz, P.G., Wilson, I.A.(2005) Protein Sci 14: 1340-1349
- PubMed: 15840835
- DOI: https://doi.org/10.1110/ps.041239305
- Primary Citation Related Structures:
1U7D, 1U7X - PubMed Abstract:
The Methanococcus jannaschii tRNA(Tyr)/TyrRS pair has been engineered to incorporate unnatural amino acids into proteins in E. coli. To reveal the structural basis for the altered specificity of mutant TyrRS for O-methyl-L-tyrosine (OMeTyr), the crystal structures for the apo wild-type and mutant M. jannaschii TyrRS were determined at 2.66 and 3.0 A, respectively, for comparison with the published structure of TyrRS complexed with tRNA(Tyr) and substrate tyrosine. A large conformational change was found for the anticodon recognition loop 257-263 of wild-type TyrRS upon tRNA binding in order to facilitate recognition of G34 of the anticodon loop through pi-stacking and hydrogen bonding interactions. Loop 133-143, which is close to the tRNA acceptor stem-binding site, also appears to be stabilized by interaction with the tRNA(Tyr). Binding of the substrate tyrosine results in subtle and cooperative movements of the side chains within the tyrosine-binding pocket. In the OMeTyr-specific mutant synthetase structure, the signature motif KMSKS loop and acceptor stem-binding loop 133-143 were surprisingly ordered in the absence of bound ATP and tRNA. The active-site mutations result in altered hydrogen bonding and steric interactions which favor binding of OMeTyr over L-tyrosine. The structure of the mutant and wild-type TyrRS now provide a basis for generating new active-site libraries to evolve synthetases specific for other unnatural amino acids.
- Department of Molecular Biology, The Scripps Research Institute, 10550 Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: