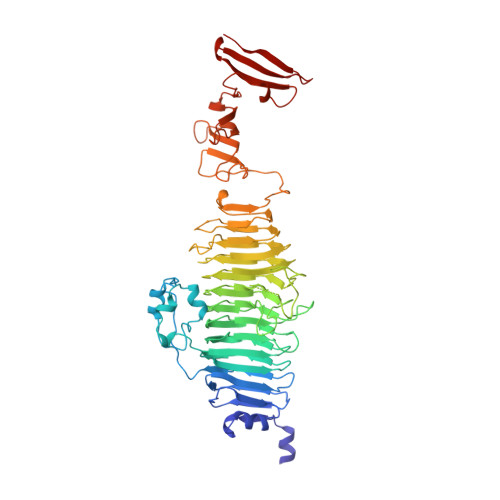
Crystal structure of phage P22 tailspike protein complexed with Salmonella sp. O-antigen receptors.
Steinbacher, S., Baxa, U., Miller, S., Weintraub, A., Seckler, R., Huber, R.(1996) Proc Natl Acad Sci U S A 93: 10584-10588
- PubMed: 8855221 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.93.20.10584
- Primary Citation Related Structures:
1TYU, 1TYV, 1TYW, 1TYX - PubMed Abstract:
The O-antigenic repeating units of lipopolysaccharides from Salmonella serogroups A, B, and D1 serve as receptors for the phage P22 tailspike protein, which also has receptor destroying endoglycosidase (endorhamnosidase) activity, integrating the functions of both hemagglutinin and neuraminidase in influenza virus. Crystal structures of the tailspike protein in complex with oligosaccharides, comprising two O-antigenic repeating units from Salmonella typhimurium, Salmonella enteritidis, and Salmonella typhi 253Ty were determined at 1.8 A resolution. The active-site topology with Asp-392, Asp-395, and Glu-359 as catalytic residues was identified. Kinetics of binding and cleavage suggest a role of the receptor destroying endorhamnosidase activity primarily for detachment of newly assembled phages.
- Abteilung Strukturforschung, Max-Planck-Institut für Biochemie, Martinsried, Germany.
Organizational Affiliation: