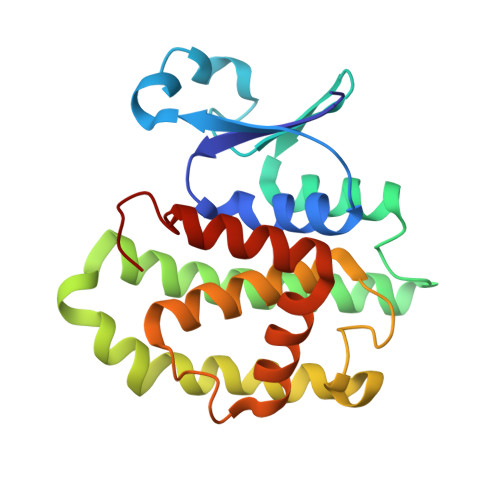
Structure of the Major Cytosolic Glutathione S-Transferase from the Parasitic Nematode Onchocerca volvulus
Perbandt, M., Hoppner, J., Betzel, C., Walter, R.D., Liebau, E.(2005) J Biological Chem 280: 12630-12636
- PubMed: 15640152
- DOI: https://doi.org/10.1074/jbc.M413551200
- Primary Citation of Related Structures:
1TU7, 1TU8 - PubMed Abstract:
Onchocerciasis is a debilitating parasitic disease caused by the filarial worm Onchocerca volvulus. Similar to other helminth parasites, O. volvulus is capable of evading the host's immune responses by a variety of defense mechanisms, including the detoxification activities of the glutathione S-transferases (GSTs). Additionally, in response to drug treatment, helminth GSTs are highly up-regulated, making them tempting targets both for chemotherapy and for vaccine development. We analyzed the three-dimensional x-ray structure of the major cytosolic GST from O. volvulus (Ov-GST2) in complex with its natural substrate glutathione and its competitive inhibitor S-hexylglutathione at 1.5 and 1.8 angstrom resolution, respectively. From the perspective of the biochemical classification, the Ov-GST2 seems to be related to pi-class GSTs. However, in comparison to other pi-class GSTs, in particular to the host's counterpart, the Ov-GST2 reveals significant and unusual differences in the sequence and overall structure. Major differences can be found in helix alpha-2, an important region for substrate recognition. Moreover, the binding site for the electrophilic co-substrate is spatially increased and more solvent-accessible. These structural alterations are responsible for different substrate specificities and will form the basis of parasite-specific structure-based drug design investigations.
- Institute of Biochemistry and Foodchemistry, Department of Biochemistry and Molecularbiology, University of Hamburg, Martin Luther King Platz 6, 20146 Hamburg, Germany. Perbandt@chemie.uni-hamburg.de
Organizational Affiliation: