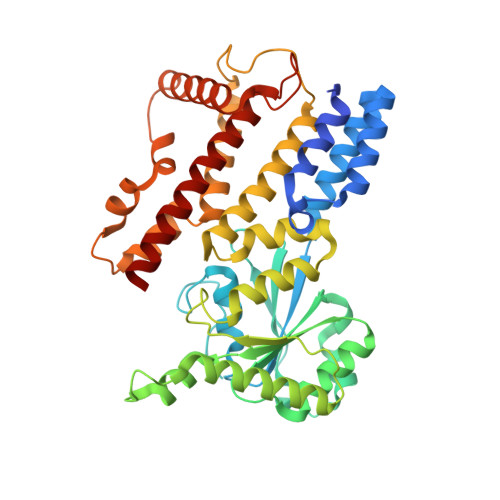
Crystal Structure of IIGP1; A Paradigm for Interferon-Inducible p47 Resistance GTPases
Ghosh, A., Uthaiah, R., Howard, J., Herrmann, C., Wolf, E.(2004) Mol Cell 15: 727-739
- PubMed: 15350217 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2004.07.017
- Primary Citation Related Structures:
1TPZ, 1TQ2, 1TQ4, 1TQ6, 1TQD - PubMed Abstract:
Interferon-inducible p47 GTPases are critical mediators of cell-autonomous resistance against several intracellular pathogens. Here we present the first crystal structure of a member of this novel GTPase family, IIGP1, in its nucleotide-free, GDP-, and GppNHp-bound form. The structure shows a Ras-like G domain between an N-terminal three-helix bundle and a complex system of C-terminal helices and loops. Sequence comparison and secondary structure prediction suggest the IIGP1 structure to be a valid model for the p47 GTPase family. The IIGP1 crystals contain a noncrystallographic dimer. We show that the dimer is required for cooperative GTP hydrolysis and GTP-dependent oligomerization of IIGP1. We also present the GDP- and GppNHp-bound monomeric structures of two dimer interface mutants. Our structures direct approaches to the analysis of the catalytic mechanism of IIGP1 and provide a coherent basis for structure-function studies aimed at elucidating the mechanistic basis of pathogen resistance caused by these enigmatic GTPases.
- Department of Structural Biology, Max Planck Institute for Molecular Physiology, Otto-Hahn-Strasse 11, 44227 Dortmund, Germany.
Organizational Affiliation: