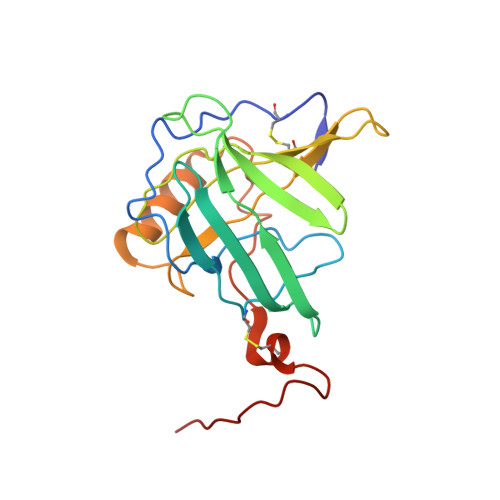
NMR structure and dynamics of the engineered fluorescein-binding lipocalin FluA reveal rigidification of beta-barrel and variable loops upon enthalpy-driven ligand binding.
Mills, J.L., Liu, G., Skerra, A., Szyperski, T.(2009) Biochemistry 48: 7411-7419
- PubMed: 19603796 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi900535j
- Primary Citation Related Structures:
1T0V - PubMed Abstract:
The NMR structure of the 21 kDa lipocalin FluA, which was previously obtained by combinatorial design, elucidates a reshaped binding site specific for the dye fluorescein resulting from 21 side chain replacements with respect to the parental lipocalin, the naturally occurring bilin-binding protein (BBP). As expected, FluA exhibits the lipocalin fold of BBP, comprising eight antiparallel beta-strands forming a beta-barrel with an alpha-helix attached to its side. Comparison of the NMR structure of free FluA with the X-ray structures of BBP.biliverdin IX(gamma) and FluA.fluorescein complexes revealed significant conformational changes in the binding pocket, which is formed by four loops at the open end of the beta-barrel as well as adjoining beta-strand segments. An "induced fit" became apparent for the side chain conformations of Arg 88 and Phe 99, which contact the bound fluorescein in the complex and undergo concerted rearrangement upon ligand binding. Moreover, slower internal motional modes of the polypeptide backbone were identified by measuring transverse (15)N backbone spin relaxation times in the rotating frame for free FluA and also for the FluA.fluorescein complex. A reduction in the level of such motions was detected upon complex formation, indicating rigidification of the protein structure and loss of conformational entropy. This hypothesis was confirmed by isothermal titration calorimetry, showing that ligand binding is enthalpy-driven, thus overcompensating for the negative entropy associated with both ligand binding per se and rigidification of the protein. Our investigation of the solution structure and dynamics as well as thermodynamics of lipocalin-ligand interaction not only provides insight into the general mechanism of small molecule accommodation in the deep and narrow cavity of this abundant class of proteins but also supports the future design of corresponding binding proteins with novel specificities, so-called "anticalins".
- Department of Chemistry, The State University of New York at Buffalo, Buffalo, New York 14260, USA.
Organizational Affiliation: