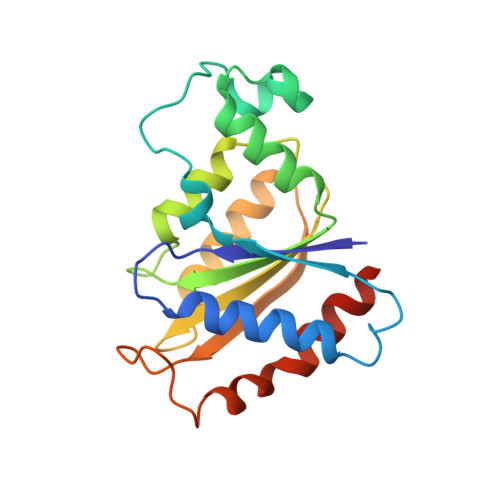
Crystal Structure and Functional Characterization of Yeast YLR011wp, an Enzyme with NAD(P)H-FMN and Ferric Iron Reductase Activities
Liger, D., Graille, M., Zhou, C.-Z., Leulliot, N., Quevillon-Cheruel, S., Blondeau, K., Janin, J., Van Tilbeurgh, H.(2004) J Biological Chem 279: 34890-34897
- PubMed: 15184374 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M405404200
- Primary Citation Related Structures:
1T0I - PubMed Abstract:
Flavodoxins are involved in a variety of electron transfer reactions that are essential for life. Although FMN-binding proteins are well characterized in prokaryotic organisms, information is scarce for eukaryotic flavodoxins. We describe the 2.0-A resolution crystal structure of the Saccharomyces cerevisiae YLR011w gene product, a predicted flavoprotein. YLR011wp indeed adopts a flavodoxin fold, binds the FMN cofactor, and self-associates as a homodimer. Despite the absence of the flavodoxin key fingerprint motif involved in FMN binding, YLR011wp binds this cofactor in a manner very analogous to classical flavodoxins. YLR011wp closest structural homologue is the homodimeric Bacillus subtilis Yhda protein (25% sequence identity) whose homodimer perfectly superimposes onto the YLR011wp one. Yhda, whose function is not documented, has 53% sequence identity with the Bacillus sp. OY1-2 azoreductase. We show that YLR011wp has an NAD(P)H-dependent FMN reductase and a strong ferricyanide reductase activity. We further demonstrate a weak but specific reductive activity on azo dyes and nitrocompounds.
- Institut de Biochimie et de Biophysique Moléculaire et Cellulaire (CNRS-Unité Mixte de Recherche (UMR) 8619), Université Paris-Sud, Bâtiment 430, 91405 Orsay, France.
Organizational Affiliation: