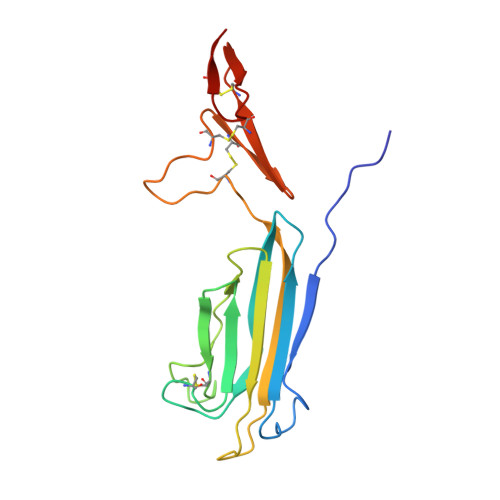
The X-ray structure of human MBL-associated protein 19 (MAp19) and its interaction site with mannan-binding lectin and L-ficolin
Gregory, L.A., Thielens, N.M., Matsushita, M., Sorensen, R., Arlaud, G.J., Fontecilla-Camps, J.C., Gaboriaud, C.(2004) J Biological Chem 279: 29391-29397
- PubMed: 15117939 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M402687200
- Primary Citation Related Structures:
1SZB - PubMed Abstract:
MAp19 is an alternative splicing product of the MASP-2 gene comprising the N-terminal CUB1-epidermal growth factor (EGF) segment of MASP-2, plus four additional residues at its C-terminal end. Like full-length MASP-2, it forms Ca(2+)-dependent complexes with mannan-binding lectin (MBL) and L-ficolin. The x-ray structure of human MAp19 was solved to a resolution of 2.5 A. It shows a head to tail homodimer held together by interactions between the CUB1 module of one monomer and the EGF module of its counterpart. A Ca(2+) ion bound to each EGF module stabilizes the dimer interfaces. A second Ca(2+) ion is bound to the distal end of each CUB1 module, through six ligands contributed by Glu(52), Asp(60), Asp(105), Ser(107), Asn(108), and a water molecule. Compared with its counterpart in human C1s, the N-terminal end of the MAp19 CUB1 module contains a 7-residue extension that forms additional inter-monomer contacts. To identify the residues involved in the interaction of MAp19 with MBL and L-ficolin, point mutants were generated and their binding ability was determined using surface plasmon resonance spectroscopy. Six mutations at Tyr(59), Asp(60), Glu(83), Asp(105), Tyr(106), and Glu(109) either strongly decreased or abolished interaction with both MBL and L-ficolin. These mutations map a common binding site for these proteins located at the distal end of each CUB1 module and stabilized by the Ca(2+) ion.
- Laboratoire de Cristallographie et Cristallogénèse des Protéines, France.
Organizational Affiliation: