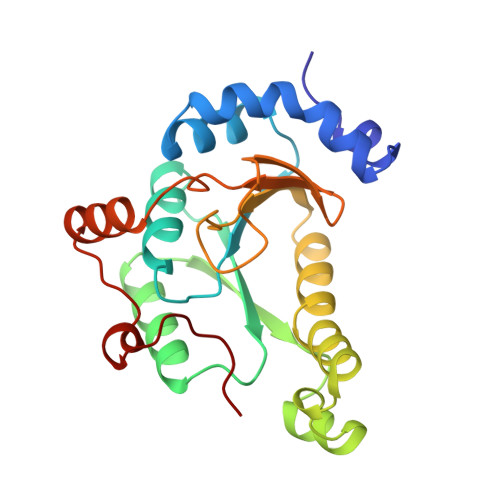
Crystal structure of phosphoadenylyl sulphate (PAPS) reductase: a new family of adenine nucleotide alpha hydrolases.
Savage, H., Montoya, G., Svensson, C., Schwenn, J.D., Sinning, I.(1997) Structure 5: 895-906
- PubMed: 9261082 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(97)00244-x
- Primary Citation Related Structures:
1SUR - PubMed Abstract:
Assimilatory sulphate reduction supplies prototrophic organisms with reduced sulphur for the biosynthesis of all sulphur-containing metabolites. This process is driven by a sequence of enzymatic steps involving phosphoadenylyl sulphate (PAPS) reductase. Thioredoxin is used as the electron donor for the reduction of PAPS to phospho-adenosine-phosphate (PAP) and sulphite. Unlike most electron-transfer reactions, there are no cofactors or prosthetic groups involved in this reduction and PAPS reductase is one of the rare examples of an enzyme that is able to store two electrons. Determination of the structure of PAPS reductase is the first step towards elucidating the biochemical details of the reduction of PAPS to sulphite. We have determined the crystal structure of PAPS reductase at 2.0 A resolution in the open, reduced form, in which a flexible loop covers the active site. The protein is active as a dimer, each monomer consisting of a central six-stranded beta sheet with alpha helices packing against each side. A highly modified version of the P loop, the fingerprint peptide of mononucleotide-binding proteins, is present in the active site of the protein, which appears to be a positively charged cleft containing a number of conserved arginine and lysine residues. Although PAPS reductase has no ATPase activity, it shows a striking similarity to the structure of the ATP pyrophosphatase (ATP PPase) domain of GMP synthetase, indicating that both enzyme families have evolved from a common ancestral nucleotide-binding fold. The sequence conservation between ATP sulphurylases, a subfamily of ATP PPases, and PAPS reductase and the similarities in both their mechanisms and folds, suggest an evolutionary link between the ATP PPases and PAPS reductases. Together with the N type ATP PPases, PAPS reductases and ATP sulphurylases are proposed to form a new family of homologous enzymes with adenine nucleotide alpha-hydrolase activity. The open, reduced form of PAPS reductase is able to bind PAPS, whereas the closed oxidized form cannot. A movement between the two monomers of the dimer may allow this switch in conformation to occur.
- European Molecular Biology Laboratory, Structural Biology Programme, Heidelberg, Germany.
Organizational Affiliation: