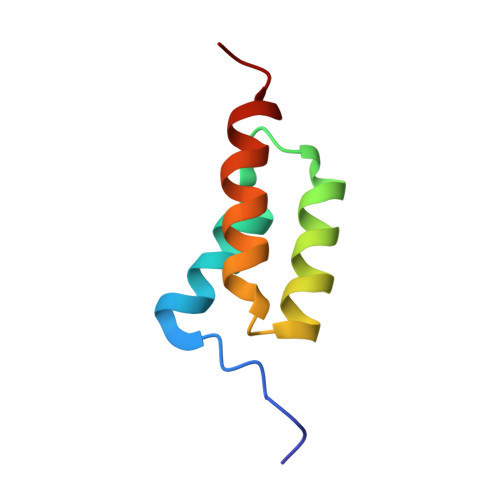
From The Cover: Testing protein-folding simulations by experiment: B domain of protein A.
Sato, S., Religa, T.L., Daggett, V., Fersht, A.R.(2004) Proc Natl Acad Sci U S A 101: 6952-6956
- PubMed: 15069202 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0401396101
- Primary Citation Related Structures:
1SS1 - PubMed Abstract:
We have assessed the published predictions of the pathway of folding of the B domain of protein A, the pathway most studied by computer simulation. We analyzed the transition state for folding of the three-helix bundle protein, by using experimental Phi values on some 70 suitable mutants. Surprisingly, the third helix, which has the most stable alpha-helical structure as a peptide fragment, is poorly formed in the transition state, especially at its C terminus. The protein folds around a nearly fully formed central helix, which is stabilized by extensive hydrophobic side chain interactions. The turn connecting the poorly structured first helix to the central helix is unstructured, but the turn connecting the central helix to the third is in the process of being formed as the N-terminal region of the third helix begins to coalesce. The transition state is inconsistent with a classical framework mechanism and is closer to nucleation-condensation. None of the published atomistic simulations are fully consistent with the experimental picture although many capture important features. There is a continuing need for combining simulation with experiment to describe folding pathways, and of continued testing to improve predictive methods.
- Medical Research Council Centre for Protein Engineering, Hills Road, Cambridge CB2 2QH, United Kingdom.
Organizational Affiliation: