Cell-free protein production and labeling protocol for NMR-based structural proteomics.
Vinarov, D.A., Lytle, B.L., Peterson, F.C., Tyler, E.M., Volkman, B.F., Markley, J.L.(2004) Nat Methods 1: 149-153
- PubMed: 15782178 Search on PubMed
- DOI: https://doi.org/10.1038/nmeth716
- Primary Citation Related Structures:
1SE9 - PubMed Abstract:
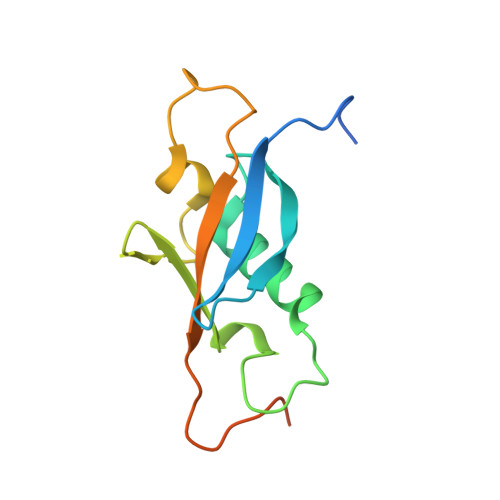
Structural proteomics requires robust, scalable methods. Here we describe a wheat germ cell-free platform for protein production that supports efficient NMR structural studies of eukaryotic proteins and offers advantages over cell-based methods. To illustrate this platform, we describe its application to a specific target (At3g01050.1) from Arabidopsis thaliana. After cloning the target gene into a specialized plasmid, we carry out a small-scale (50 mul) in vitro sequential transcription and translation trial to ascertain the level of protein production and solubility. Next, we prepare mRNA for use in a 4-ml semicontinuous cell-free translation reaction to incorporate (15)N-labeled amino acids into a protein sample that we purify and test for suitability for NMR structural analysis. We then repeat the cell-free approach with (13)C,(15)N-labeled amino acids to prepare a doubly labeled sample. The three-dimensional (3D) structure of At3g01050.1 shows that this protein is an unusual member of the beta-grasp protein family.
- Center for Eukaryotic Structural Genomics, University of Wisconsin-Madison, 433 Babcock Drive, Madison, Wisconsin 53706, USA.
Organizational Affiliation: