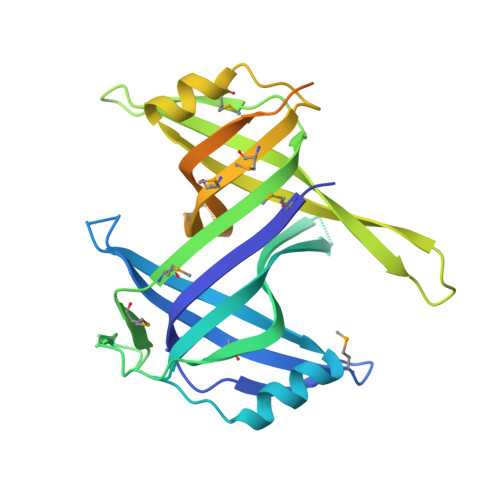
Crystal structure of the Deinococcus radiodurans single-stranded DNA-binding protein suggests a mechanism for coping with DNA damage.
Bernstein, D.A., Eggington, J.M., Killoran, M.P., Misic, A.M., Cox, M.M., Keck, J.L.(2004) Proc Natl Acad Sci U S A 101: 8575-8580
- PubMed: 15159541 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0401331101
- Primary Citation Related Structures:
1SE8 - PubMed Abstract:
Single-stranded DNA (ssDNA)-binding (SSB) proteins are uniformly required to bind and protect single-stranded intermediates in DNA metabolic pathways. All bacterial and eukaryotic SSB proteins studied to date oligomerize to assemble four copies of a conserved domain, called an oligonucleotide/oligosaccharide-binding (OB) fold, that cooperate in nonspecific ssDNA binding. The vast majority of bacterial SSB family members function as homotetramers, with each monomer contributing a single OB fold. However, SSB proteins from the Deinococcus-Thermus genera are exceptions to this rule, because they contain two OB folds per monomer. To investigate the structural consequences of this unusual arrangement, we have determined a 1.8-A-resolution x-ray structure of Deinococcus radiodurans SSB. The structure shows that D. radiodurans SSB comprises two OB domains linked by a beta-hairpin motif. The protein assembles a four-OB-fold arrangement by means of symmetric dimerization. In contrast to homotetrameric SSB proteins, asymmetry exists between the two OB folds of D. radiodurans SSB because of sequence differences between the domains. These differences appear to reflect specialized roles that have evolved for each domain. Extensive crystallographic contacts link D. radiodurans SSB dimers in an arrangement that has important implications for higher-order structures of the protein bound to ssDNA. This assembly utilizes the N-terminal OB domain and the beta-hairpin structure that is unique to Deinococcus and Thermus species SSB proteins. We hypothesize that differences between D. radiodurans SSB and homotetrameric bacterial SSB proteins may confer a selective advantage to D. radiodurans cells that aids viability in environments that challenge genomic stability.
- Department of Biomolecular Chemistry, 550 Medical Science Center, 1300 University Avenue, University of Wisconsin Medical School, Madison, WI 53706-1532, USA.
Organizational Affiliation: