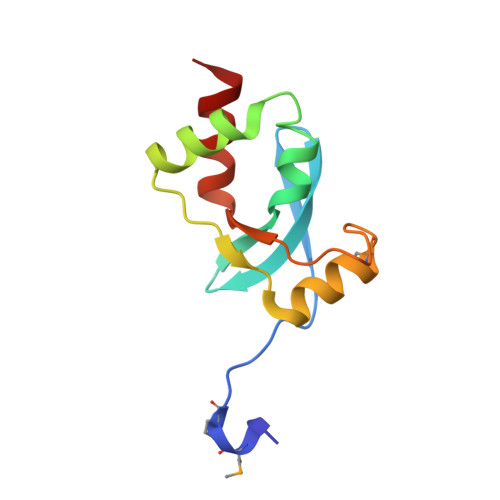
Crystal Structure of the Dachshund Homology Domain of human SKI
Wilson, J.J., Malakhova, M., Zhang, R., Joachimiak, A., Hegde, R.S.(2004) Structure 12: 785-792
- PubMed: 15130471 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.02.035
- Primary Citation Related Structures:
1SBX - PubMed Abstract:
The nuclear protooncoprotein SKI negatively regulates transforming growth factor-beta (TGF-beta) signaling in cell growth and differentiation. It directly interacts with the Smads and, by various mechanisms, represses the transcription of TGF-beta-responsive genes. SKI is a multidomain protein that includes a domain bearing high sequence similarity with the retinal determination protein Dachshund (the Dachshund homology domain, DHD). The SKI-DHD has been implicated in SMAD-2/3, N-CoR, SKIP, and PML-RARalpha binding. The 1.65 A crystal structure of the Dachshund homology domain of human SKI is reported here. The SKI-DHD adopts a mixed alpha/beta structure which includes features found in the forkhead/winged-helix family of DNA binding proteins, although SKI-DHD is not a DNA binding domain. Residues that form a contiguous surface patch on SKI-DHD are conserved within the Ski/Sno family and with Dachshund, suggesting that this domain may mediate intermolecular interactions common to these proteins.
- Division of Developmental Biology, Cincinnati Children's Hospital Medical Center and University of Cincinnati College of Medicine, 3333 Burnet Avenue, Cincinnati, OH 45229, USA.
Organizational Affiliation: