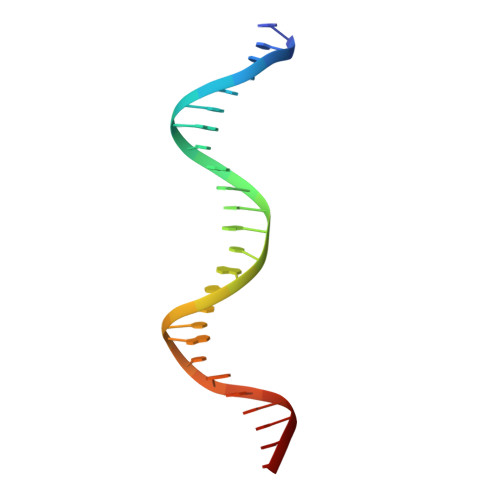
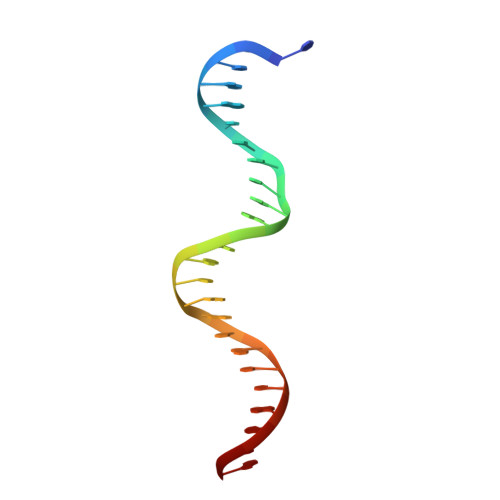
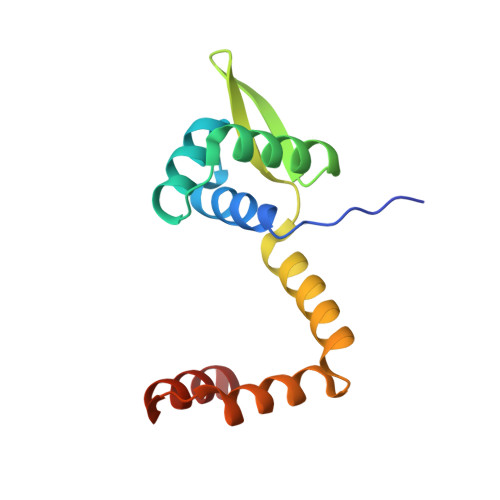
On the transcriptional regulation of methicillin resistance: MecI repressor in complex with its operator
Garcia-Castellanos, R., Mallorqui-Fernandez, G., Marrero, A., Potempa, J., Coll, M., Gomis-Ruth, F.X.(2004) J Biological Chem 279: 17888-17896
- PubMed: 14960592 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M313123200
- Primary Citation Related Structures:
1SAX - PubMed Abstract:
Bacterial resistance to antibiotics poses a serious worldwide public health problem due to the high morbidity and mortality caused by infectious diseases. Most hospital-onset infections are associated with methicillin-resistant Staphylococcus aureus (MRSA) strains that have acquired multiple drug resistance to beta-lactam antibiotics. In a response to antimicrobial stress, nearly all clinical MRSA isolates produce beta-lactamase (BlaZ) and a penicillin-binding protein with low affinity for beta-lactam antibiotics (PBP2a, also known as PBP2' or MecA). Both effectors are regulated by homologous signal transduction systems consisting of a sensor/transducer and a transcriptional repressor. MecI (methicillin repressor) blocks mecA but also blaZ transcription and that of itself and the co-transcribed sensor/transducer. The structure of MecI in complex with a cognate operator double-stranded DNA reveals a homodimeric arrangement with a novel C-terminal spiral staircase dimerization domain responsible for dimer integrity. Each protomer interacts with the DNA major groove through a winged helix DNA-binding domain and specifically recognizes the nucleotide sequence 5'-Gua-Thy-Ade-X-Thy-3'. This results in an unusual convex bending of the DNA helix. The structure of this first molecular determinant of methicillin resistance in complex with its target DNA provides insights into its regulatory mechanism and paves the way for new antimicrobial strategies against MRSA.
- Institut de Biologia Molecular de Barcelona, CID-CSIC C/Jordi Girona, 18-26, 08034 Barcelona, Spain.
Organizational Affiliation: