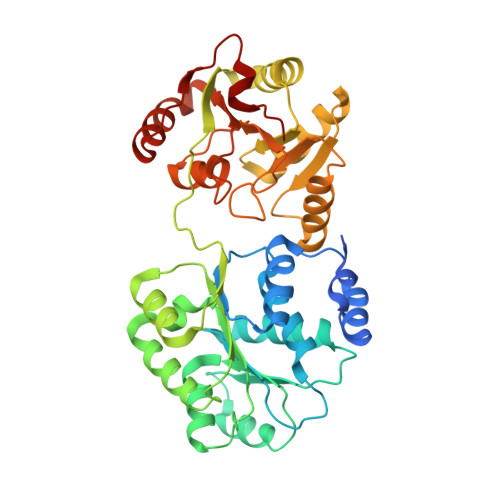
Crystal structure and functional analysis of DEAD-box protein Dhh1p.
Cheng, Z., Coller, J., Parker, R., Song, H.(2005) RNA 11: 1258-1270
- PubMed: 15987810
- DOI: https://doi.org/10.1261/rna.2920905
- Primary Citation of Related Structures:
1S2M - PubMed Abstract:
The control of mRNA translation and degradation are critical for proper gene expression. A key regulator of both translation and degradation is Dhh1p, which is a DEAD-box protein, and functions both to repress translation and enhance decapping. We describe the crystal structure of the N- and C-terminal truncated Dhh1p (tDhh1p) determined at 2.1 A resolution. This reveals that, like other DEAD-box proteins, tDhh1p contains two RecA-like domains, although with a unique arrangement. In contrast to eIF4A and mjDEAD, in which no motif interactions exist, in Dhh1p, motif V interacts with motif I and the Q-motif, thereby linking the two domains together. Electrostatic potential mapping combined with mutagenesis reveals that motifs I, V, and VI are involved in RNA binding. In addition, trypsin digestion of tDhh1p suggests that ATP binding enhances an RNA-induced conformational change. Interestingly, some mutations located in the conserved motifs and at the interface between the two Dhh1 domains confer dominant negative phenotypes in vivo and disrupt the conformational switch in vitro. This suggests that this conformational change is required in Dhh1 function and identifies key residues involved in that transition.
- Laboratory of Macromolecular Structure, Institute of Molecular and Cell Biology, Proteos, Singapore 138673.
Organizational Affiliation: