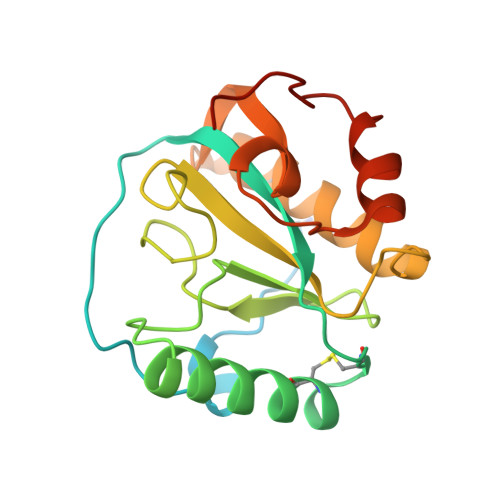
A Drosophila pattern recognition receptor contains a peptidoglycan docking groove and unusual l,d-carboxypeptidase activity.
Chang, C.-I., Pili-Floury, S., Herve, M., Parquet, C., Chelliah, Y., Lemaitre, B., Mengin-Lecreulx, D., Deisenhofer, J.(2004) PLoS Biol 2: 1293-1302
- PubMed: 15361936
- DOI: https://doi.org/10.1371/journal.pbio.0020277
- Primary Citation Related Structures:
1S2J - PubMed Abstract:
The Drosophila peptidoglycan recognition protein SA (PGRP-SA) is critically involved in sensing bacterial infection and activating the Toll signaling pathway, which induces the expression of specific antimicrobial peptide genes. We have determined the crystal structure of PGRP-SA to 2.2-A resolution and analyzed its peptidoglycan (PG) recognition and signaling activities. We found an extended surface groove in the structure of PGRP-SA, lined with residues that are highly diverse among different PGRPs. Mutational analysis identified it as a PG docking groove required for Toll signaling and showed that residue Ser158 is essential for both PG binding and Toll activation. Contrary to the general belief that PGRP-SA has lost enzyme function and serves primarily for PG sensing, we found that it possesses an intrinsic L,D-carboxypeptidase activity for diaminopimelic acid-type tetrapeptide PG fragments but not lysine-type PG fragments, and that Ser158 and His42 may participate in the hydrolytic activity. As L,D-configured peptide bonds exist only in prokaryotes, this work reveals a rare enzymatic activity in a eukaryotic protein known for sensing bacteria and provides a possible explanation of how PGRP-SA mediates Toll activation specifically in response to lysine-type PG.
- Howard Hughes Medical Institute and Department of Biochemistry, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
Organizational Affiliation: