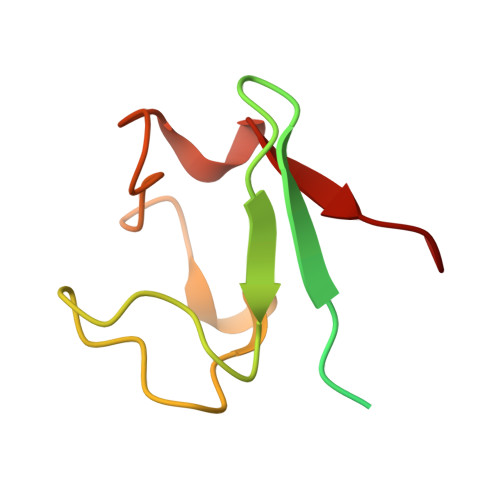
Solution structure of the two-iron rubredoxin of Pseudomonas oleovorans determined by NMR spectroscopy and solution X-ray scattering and interactions with rubredoxin reductase.
Perry, A., Tambyrajah, W., Grossmann, J.G., Lian, L.Y., Scrutton, N.S.(2004) Biochemistry 43: 3167-3182
- PubMed: 15023067 Search on PubMed
- DOI: https://doi.org/10.1021/bi035817u
- Primary Citation Related Structures:
1S24 - PubMed Abstract:
Here we provide insights into the molecular structure of the two-iron 19-kDa rubredoxin (AlkG) of Pseudomonas oleovorans using solution-state nuclear magnetic resonance (NMR) and small-angle X-ray scattering studies. Sequence alignment and biochemical studies have suggested that AlkG comprises two rubredoxin folds connected by a linker region of approximately 70 amino acid residues. The C-terminal domain (C-Rb) of this unusual rubredoxin, together with approximately 35 amino acid residues of the predicted linker region, was expressed in Escherichia coli, purified in the one-iron form and the structure of the cadmium-substituted form determined at high-resolution by NMR spectroscopy. The structure shows that the C-Rb domain is similar in fold to the conventional one-iron rubredoxins from other organisms, whereas the linker region does not have any discernible structure. This tandem "flexible-folded" structure of the polypeptide chain derived for the C-Rb protein was confirmed using solution X-ray scattering methods. X-ray scattering studies of AlkG indicated that the 70-amino acid residue linker forms a structured, yet mobile, polypeptide segment connecting the globular N- and C-terminal domains. The X-ray scattering studies also showed that the N-terminal domain (N-Rb) has a molecular conformation similar to that of C-Rb. The restored molecular shape indicates that the folded N-Rb and C-Rb domains of AlkG are noticeably separated, suggesting some domain movement on complex formation with rubredoxin reductase to allow interdomain electron transfer between the metal centers in AlkG. This study demonstrates the advantage of combining X-ray scattering and NMR methods in structural studies of dynamic, multidomain proteins that are not suited to crystallographic analysis. The study forms a structural foundation for functional studies of the interaction and electron-transfer reactions of AlkG with rubredoxin reductase, also reported herein.
- Department of Biochemistry and Centre for Chemical Biology, University of Leicester, UK.
Organizational Affiliation: