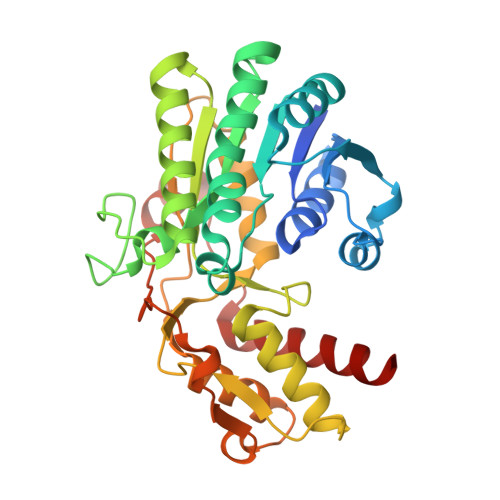
Crystal structure of a tetrameric GDP-D-mannose 4,6-dehydratase from a bacterial GDP-D-rhamnose biosynthetic pathway.
Webb, N.A., Mulichak, A.M., Lam, J.S., Rocchetta, H.L., Garavito, R.M.(2004) Protein Sci 13: 529-539
- PubMed: 14739333 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.03393904
- Primary Citation Related Structures:
1RPN - PubMed Abstract:
d-Rhamnose is a rare 6-deoxy monosaccharide primarily found in the lipopolysaccharide of pathogenic bacteria, where it is involved in host-bacterium interactions and the establishment of infection. The biosynthesis of d-rhamnose proceeds through the conversion of GDP-d-mannose by GDP-d-mannose 4,6-dehydratase (GMD) to GDP-4-keto-6-deoxymannose, which is subsequently reduced to GDP-d-rhamnose by a reductase. We have determined the crystal structure of GMD from Pseudomonas aeruginosa in complex with NADPH and GDP. GMD belongs to the NDP-sugar modifying subfamily of the short-chain dehydrogenase/reductase (SDR) enzymes, all of which exhibit bidomain structures and a conserved catalytic triad (Tyr-XXX-Lys and Ser/Thr). Although most members of this enzyme subfamily display homodimeric structures, this bacterial GMD forms a tetramer in the same fashion as the plant MUR1 from Arabidopsis thaliana. The cofactor binding sites are adjoined across the tetramer interface, which brings the adenosyl phosphate moieties of the adjacent NADPH molecules to within 7 A of each other. A short peptide segment (Arg35-Arg43) stretches into the neighboring monomer, making not only protein-protein interactions but also hydrogen bonding interactions with the neighboring cofactor. The interface hydrogen bonds made by the Arg35-Arg43 segment are generally conserved in GMD and MUR1, and the interacting residues are highly conserved among the sequences of bacterial and eukaryotic GMDs. Outside of the Arg35-Arg43 segment, residues involved in tetrameric contacts are also quite conserved across different species. These observations suggest that a tetramer is the preferred, and perhaps functionally relevant, oligomeric state for most bacterial and eukaryotic GMDs.
- Department of Biochemistry and Molecular Biology, Michigan State University, East Lansing, MI 48824-1319, USA.
Organizational Affiliation: