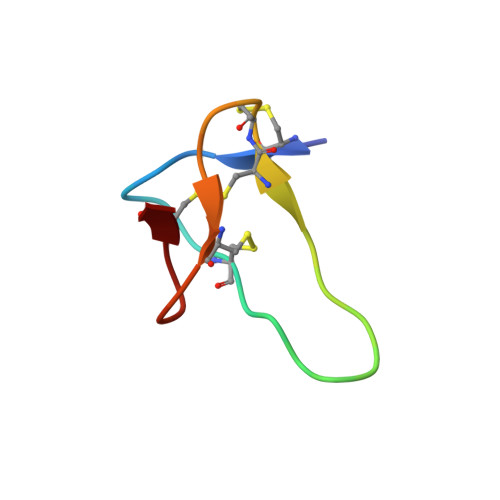
Structures of muO-conotoxins from Conus marmoreus. Inhibitors of tetrodotoxin (TTX)-sensitive and TTX-resistant sodium channels in mammalian sensory neurons
Daly, N.L., Ekberg, J.A., Thomas, L., Adams, D.J., Lewis, R.J., Craik, D.J.(2004) J Biological Chem 279: 25774-25782
- PubMed: 15044438 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M313002200
- Primary Citation Related Structures:
1RMK - PubMed Abstract:
The microO-conotoxins are an intriguing class of conotoxins targeting various voltage-dependent sodium channels and molluscan calcium channels. In the current study, we have shown MrVIA and MrVIB to be the first known peptidic inhibitors of the transient tetrodotoxin-resistant (TTX-R) Na(+) current in rat dorsal root ganglion neurons, in addition to inhibiting tetrodotoxin-sensitive Na(+) currents. Human TTX-R sodium channels are a therapeutic target for indications such as pain, highlighting the importance of the microO-conotoxins as potential leads for drug development. Furthermore, we have used NMR spectroscopy to provide the first structural information on this class of conotoxins. MrVIA and MrVIB are hydrophobic peptides that aggregate in aqueous solution but were solubilized in 50% acetonitrile/water. The three-dimensional structure of MrVIB consists of a small beta-sheet and a cystine knot arrangement of the three-disulfide bonds. It contains four backbone "loops" between successive cysteine residues that are exposed to the solvent to varying degrees. The largest of these, loop 2, is the most disordered part of the molecule, most likely due to flexibility in solution. This disorder is the most striking difference between the structures of MrVIB and the known delta- and omega-conotoxins, which along with the microO-conotoxins are members of the O superfamily. Loop 2 of omega-conotoxins has previously been shown to contain residues critical for binding to voltage-gated calcium channels, and it is interesting to speculate that the flexibility observed in MrVIB may accommodate binding to both sodium and molluscan calcium channels.
- Institute for Molecular Bioscience and School of Biomedical Sciences, University of Queensland, Brisbane, 4072 Queensland, Australia.
Organizational Affiliation: