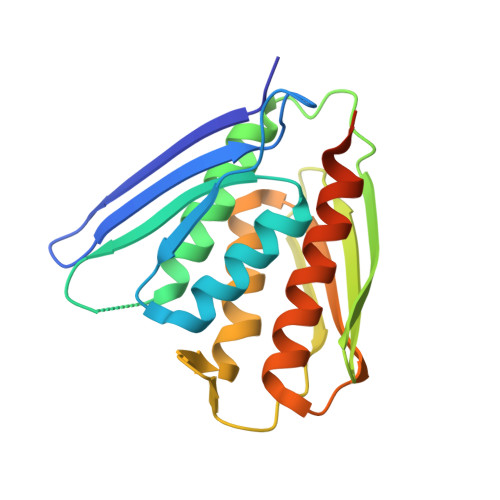
Crystal structure of imidazole glycerol-phosphate dehydratase: duplication of an unusual fold
Sinha, S.C., Chaudhuri, B.N., Burgner, J.W., Yakovleva, G., Davisson, V.J., Smith, J.L.(2004) J Biological Chem 279: 15491-15498
- PubMed: 14724278 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M312733200
- Primary Citation Related Structures:
1RHY - PubMed Abstract:
Imidazole glycerol-phosphate dehydratase (IGPD) catalyzes the sixth step of histidine biosynthesis. The enzyme is of fundamental biochemical interest, because it catalyzes removal of a non-acidic hydrogen atom in the dehydration reaction. It is also a potential target for development of herbicides. IGPD is a metalloenzyme in which transition metals induce aggregation and are required for catalysis. Addition of 1 equivalent of Mn(2+)/subunit is shown by analytical ultracentrifugation to induce the formation of 24-mers from trimeric IGPD. Two histidine-rich motifs may participate in metal binding and aggregation. The 2.3-A crystal structure of metal-free trimeric IGPD from the fungus Filobasidiella neoformans reveals a novel fold containing an internal repeat, apparently the result of gene duplication. The 95-residue alpha/beta half-domain occurs in a few other proteins, including the GHMP kinase superfamily (galacto-homoserine-mevalonate-phosphomevalonate), but duplication to form a compact domain has not been seen elsewhere. Conserved residues cluster at two types of sites in the trimer, each site containing a conserved histidine-rich motif. A model is proposed for the intact, active 24-mer in which all highly conserved residues, including the histidine-rich motifs in both the N- and C-terminal halves of the polypeptide, cluster at a common site between trimers. This site is a candidate for the active site and also for metal binding leading to aggregation of trimers. The structure provides a basis for further studies of enzyme function and mechanism and for development of more potent and specific herbicides.
- Department of Biological Sciences, Purdue University, West Lafayette, Indiana 47907, USA.
Organizational Affiliation: