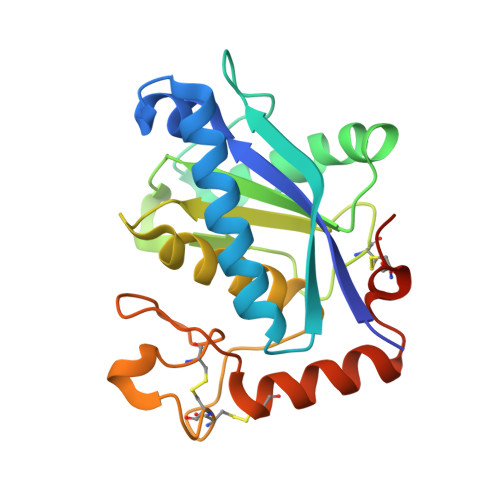
Crystal structre of the catalytic domain of human ADAM33
Orth, P., Reichert, P., Wang, W., Prosise, W.W., Yarosh-Tomaine, T., Hammond, G., Xiao, L., Mirza, U.A., Zou, J., Strickland, C., Taremi, S.S., Le, H.V., Madison, V.(2004) J Mol Biology 335: 129-137
- PubMed: 14659745 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2003.10.037
- Primary Citation Related Structures:
1R54, 1R55 - PubMed Abstract:
Adam33 is a putative asthma susceptibility gene encoding for a membrane-anchored metalloprotease belonging to the ADAM family. The ADAMs (a disintegrin and metalloprotease) are a family of glycoproteins implicated in cell-cell interactions, cell fusion, and cell signaling. We have determined the crystal structure of the Adam33 catalytic domain in complex with the inhibitor marimastat and the inhibitor-free form. The structures reveal the polypeptide fold and active site environment resembling that of other metalloproteases. The substrate-binding site contains unique features that allow the structure-based design of specific inhibitors of this enzyme.
- Schering-Plough Research Institute, 2015 Galloping Hill Rd, Kenilworth, NJ 07033, USA. peter.orth@spcorp.com
Organizational Affiliation: