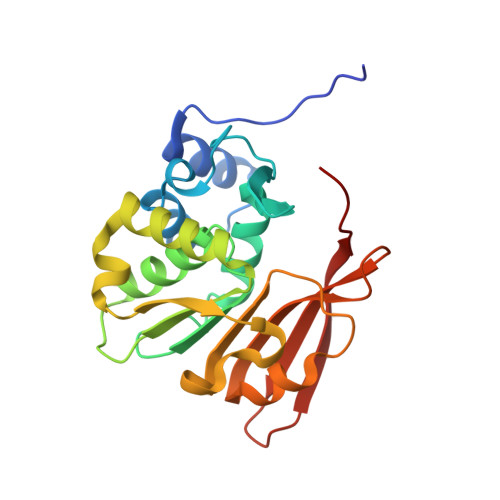
Catalytic implications from the Drosophila protein L-isoaspartyl methyltransferase structure and site-directed mutagenesis.
Bennett, E.J., Bjerregaard, J., Knapp, J.E., Chavous, D.A., Friedman, A.M., Royer Jr., W.E., O'Connor, C.M.(2003) Biochemistry 42: 12844-12853
- PubMed: 14596598
- DOI: https://doi.org/10.1021/bi034891+
- Primary Citation Related Structures:
1R18 - PubMed Abstract:
Protein L-isoaspartyl methyltransferases (PIMT; EC 2.1.1.77) catalyze the S-adenosylmethionine-dependent methylation of L-isoaspartyl residues that arise spontaneously in proteins with age, thereby initiating a repair process that restores the normal backbone configuration to the damaged polypeptide. In Drosophila melanogaster, overexpression of PIMT in transgenic flies extends the normal life span, suggesting that protein damage can be a limiting factor in longevity. To understand structural features of the Drosophila PIMT (dPIMT) important for catalysis, the crystal structure of dPIMT was determined at a resolution of 2.2 A, and site-directed mutagenesis was used to identify the role of Ser-60 in catalysis. The core structure of dPIMT is similar to the modified nucleotide-binding fold observed in PIMTs from extreme thermophiles and humans. A striking difference of the dPIMT structure is the rotation of the C-terminal residues by 90 degrees relative to the homologous structures. Effectively, this displacement generates a more open conformation that allows greater solvent access to S-adenosylhomocysteine, which is almost completely buried in other PIMT structures. The enzyme may alternate between the open conformation found for dPIMT and the more closed conformations described for other PIMTs during its catalytic cycle, thereby allowing the exchange of substrates and products. Catalysis by dPIMT requires the side chain of the conserved, active site residue Ser-60, since substitution of this residue with Thr, Gln, or Ala reduces or abolishes the methylation of both protein and isoaspartyl peptide substrates.
- Biology Department, Boston College, Chestnut Hill, Massachusetts 02467, USA.
Organizational Affiliation: