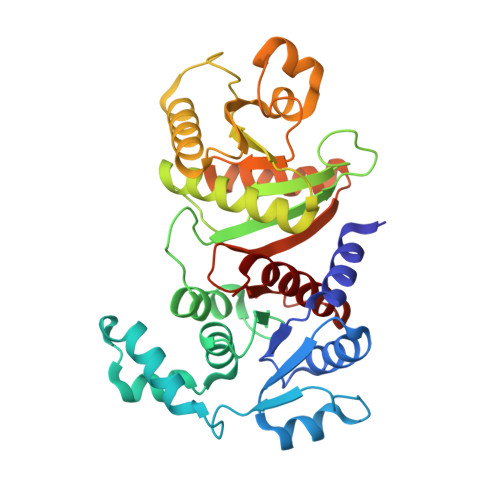
Crystal structure of phosphotransacetylase from the methanogenic archaeon Methanosarcina thermophila.
Iyer, P.P., Lawrence, S.H., Luther, K.B., Rajashankar, K.R., Yennawar, H.P., Ferry, J.G., Schindelin, H.(2004) Structure 12: 559-567
- PubMed: 15062079
- DOI: https://doi.org/10.1016/j.str.2004.03.007
- Primary Citation of Related Structures:
1QZT - PubMed Abstract:
Phosphotransacetylase (Pta) [EC 2.3.1.8] is ubiquitous in the carbon assimilation and energy-yielding pathways in anaerobic prokaryotes where it catalyzes the reversible transfer of the acetyl group from acetyl phosphate to CoA forming acetyl CoA and inorganic phosphate. The crystal structure of Pta from the methane-producing archaeon Methanosarcina thermophila, representing the first crystal structure of any Pta, was determined by multiwavelength anomalous diffraction at 2.7 A resolution. In solution and in the crystal, the enzyme forms a homodimer. Each monomer consists of two alpha/beta domains with a cleft along the domain boundary, which presumably contains the substrate binding sites. Comparison of the four monomers present in the asymmetric unit indicates substantial variations in the relative orientation of the two domains and the structure of the putative active site cleft. A search for structural homologs revealed the NADP(+)-dependent isocitrate and isopropylmalate dehydrogenases as the only homologs with a similar two-domain architecture.
- Department of Biochemistry and Molecular Biology, The Pennsylvania State University, University Park, PA 16802, USA.
Organizational Affiliation: