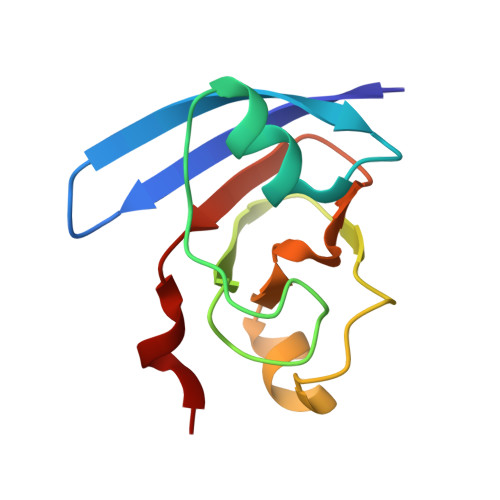
Refined X-ray structures of the oxidized, at 1.3 A, and reduced, at 1.17 A, [2Fe-2S] ferredoxin from the cyanobacterium Anabaena PCC7119 show redox-linked conformational changes.
Morales, R., Charon, M.H., Hudry-Clergeon, G., Petillot, Y., Norager, S., Medina, M., Frey, M.(1999) Biochemistry 38: 15764-15773
- PubMed: 10625442 Search on PubMed
- DOI: https://doi.org/10.1021/bi991578s
- Primary Citation Related Structures:
1CZP, 1QT9 - PubMed Abstract:
The chemical sequence of the [2Fe-2S] ferredoxin from the cyanobacterium AnabaenaPCC7119 (Fd7119) and its high-resolution X-ray structures in the oxidized and reduced states have been determined. The Fd7119 sequence is identical to that of the ferredoxin from the PCC7120 strain (Fd7120). X-ray diffraction data were collected at 100 K with an oxidized trigonal Fd7119 crystal, at 1.3 A resolution, and with an orthorhombic crystal, previously reduced with dithionite and flash frozen under anaerobic conditions, at 1.17 A resolution. The two molecular models were determined by molecular replacement with the [2Fe-2S] ferredoxin from the strain PCC7120 (Rypniewski, W. R., Breiter, D. R., Benning, M. M., Wesenberg, G., Oh, B.-H., Markley, J. L., Rayment, I., and Holden, H. M. (1991) Biochemistry 30, 4126-4131.) The final R-factors are 0. 140 (for the reduced crystal) and 0.138 (for the oxidized crystal). The [2Fe-2S] cluster appears as a significantly distorted lozenge in the reduced and oxidized redox states. The major conformational difference between the two redox forms concerns the peptide bond linking Cys46 and Ser47 which points its carbonyl oxygen away from the [2Fe-2S] cluster ("CO out") in the reduced molecule and toward it ("CO in") in the oxidized one. The "CO out" conformation could be the signature of the reduction of the iron atom Fe1, which is close to the molecular surface. Superposition of the three crystallographically independent molecules shows that the putative recognition site with the physiological partner (FNR) involves charged, hydrophobic residues and invariant water molecules.
- Laboratoire de Cristallographie et de Cristallogénèse des Protéines, Laboratoire d'Enzymologie Moléculaire, and Laboratoire de Spectrométrie de Masse, Institut de Biologie Structurale J.P. Ebel, CEA-CNRS, 41 rue Jules Horowitz, France.
Organizational Affiliation: