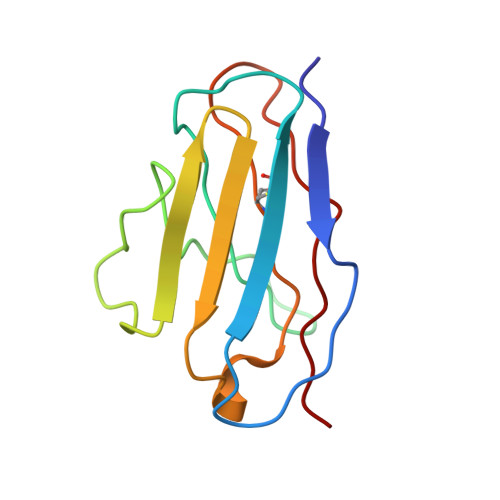
Molecular structure of the amyloid-forming protein kappa I Bre.
Steinrauf, L.K., Chiang, M.Y., Shiuan, D.(1999) J Biochem 125: 422-429
- PubMed: 9990143 Search on PubMed
- DOI: https://doi.org/10.1093/oxfordjournals.jbchem.a022303
- Primary Citation Related Structures:
1QP1 - PubMed Abstract:
The molecular structure of the amyloid-forming Bence-Jones protein kappa I Bre has been determined by X-ray crystallography at 2.0 A resolution. The fragment from the kappa chain of immunoprotein contains 107 amino acid residues, and polymerizes in the crystal form into a giant helical spiral, surrounding a cylinder of water 50 A in diameter with a repeat of 77.56 A, containing 12 kappa molecules, plus another 12 molecules from neighboring parallel spirals. The resulting structure has many features which have been found or suggested from studies on the protein fibrils found in amyloid deposits. From the results of the X-ray crystal structure a hypothesis is presented for the structure and formation of the amyloid fibril.
- Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, Indianapolis, Indiana, USA. lsteinra@www.ora.nsysu.edu.tw
Organizational Affiliation: