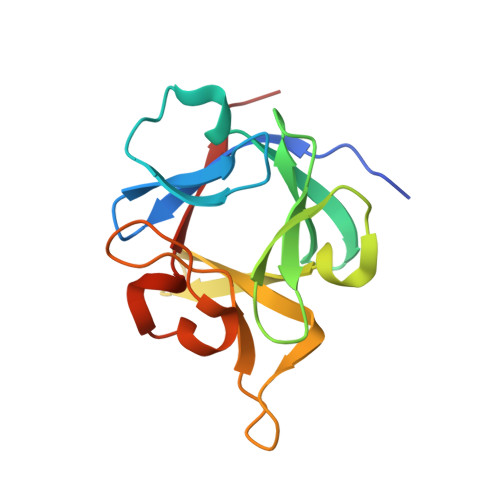
Fibroblast growth factor (FGF) homologous factors share structural but not functional homology with FGFs
Olsen, S.K., Garbi, M., Zampieri, N., Eliseenkova, A.V., Ornitz, D.M., Goldfarb, M., Mohammadi, M.(2003) J Biological Chem 278: 34226-34236
- PubMed: 12815063 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M303183200
- Primary Citation Related Structures:
1Q1U - PubMed Abstract:
Fibroblast growth factors (FGFs) interact with heparan sulfate glycosaminoglycans and the extracellular domains of FGF cell surface receptors (FGFRs) to trigger receptor activation and biological responses. FGF homologous factors (FHF1-FHF4; also known as FGF11-FGF14) are related to FGFs by substantial sequence homology, yet their only documented interactions are with an intracellular kinase scaffold protein, islet brain-2 (IB2) and with voltage-gated sodium channels. In this report, we show that recombinant FHFs can bind heparin with high affinity like classical FGFs yet fail to activate any of the seven principal FGFRs. Instead, we demonstrate that FHFs bind IB2 directly, furthering the contention that FHFs and FGFs elicit their biological effects by binding to different protein partners. To understand the molecular basis for this differential target binding specificity, we elucidated the crystal structure of FHF1b to 1.7-A resolution. The FHF1b core domain assumes a beta-trefoil fold consisting of 12 antiparallel beta strands (beta 1 through beta 12). The FHF1b beta-trefoil core is remarkably similar to that of classical FGFs and exhibits an FGF-characteristic heparin-binding surface as attested to by the number of bound sulfate ions. Using molecular modeling and structure-based mutational analysis, we identified two surface residues, Arg52 in the beta 4-beta 5 loop and Val95 in the beta 9 strand of FHF1b that are required for the interaction of FHF1b with IB2. These two residues are unique to FHFs, and mutations of the corresponding residues of FGF1 to Arg and Val diminish the capacity of FGF1 to activate FGFRs, suggesting that these two FHF residues contribute to the inability of FHFs to activate FGFRs. Hence, FHFs and FGFs bear striking structural similarity but have diverged to direct related surfaces toward interaction with distinct protein targets.
- Department of Pharmacology, New York University School of Medicine, New York, New York 10016, USA.
Organizational Affiliation: