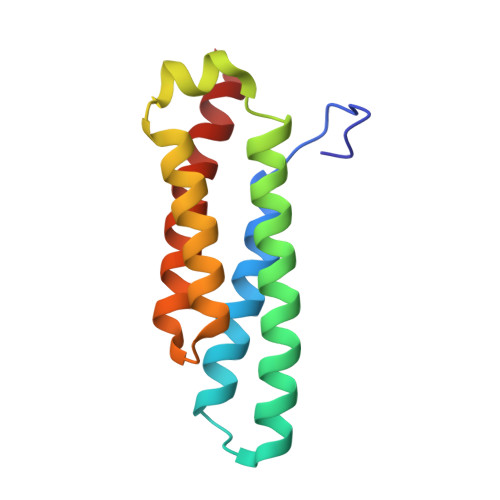
Crystal structure of nickel-containing superoxide dismutase reveals another type of active site
Wuerges, J., Lee, J.-W., Yim, Y.-I., Yim, H.-S., Kang, S.-O., Djinovic Carugo, K.(2004) Proc Natl Acad Sci U S A 101: 8569-8574
- PubMed: 15173586
- DOI: https://doi.org/10.1073/pnas.0308514101
- Primary Citation Related Structures:
1Q0D, 1Q0F, 1Q0G, 1Q0K, 1Q0M - PubMed Abstract:
Superoxide dismutases (SODs, EC 1.15.1.1) are ubiquitous enzymes that efficiently catalyze the dismutation of superoxide radical anions to protect biological molecules from oxidative damage. The crystal structure of nickel-containing SOD (NiSOD) from Streptomyces seoulensis was determined for the resting, x-ray-reduced, and thiosulfate-reduced enzyme state. NiSOD is a homohexamer consisting of four-helix-bundle subunits. The catalytic center resides in the N-terminal active-site loop, where a Ni(III) ion is coordinated by the amino group of His-1, the amide group of Cys-2, two thiolate groups of Cys-2 and Cys-6, and the imidazolate of His-1 as axial ligand that is lost in the chemically reduced state as well as after x-ray-induced reduction. This structure represents a third class of SODs concerning the catalytic metal species, subunit structure, and oligomeric organization. It adds a member to the small number of Ni-metalloenzymes and contributes with its Ni(III) active site to the general understanding of Ni-related biochemistry. NiSOD is shown to occur also in bacteria other than Streptomyces and is predicted to be present in some cyanobacteria.
- International School for Advanced Studies, Via Beirut 2-4, I-34014 Trieste, Italy.
Organizational Affiliation: