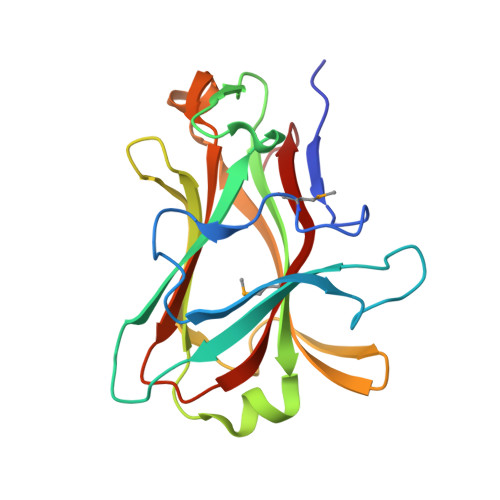
High-resolution crystal structures of Caldicellulosiruptor strain Rt8B.4 carbohydrate-binding module CBM27-1 and its complex with mannohexaose.
Roske, Y., Sunna, A., Pfeil, W., Heinemann, U.(2004) J Mol Biology 340: 543-554
- PubMed: 15210353 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.04.072
- Primary Citation Related Structures:
1PMH, 1PMJ - PubMed Abstract:
Carbohydrate-binding modules (CBMs) are the most common non-catalytic modules associated with enzymes active in plant cell-wall hydrolysis. Despite the large number of putative CBMs being identified by amino acid sequence alignments, only few representatives have been experimentally shown to have a carbohydrate-binding function. Caldicellulosiruptor strain Rt8B.4 Man26 is a thermostable modular glycoside hydrolase beta-mannanase which contains two non-catalytic modules in tandem at its N terminus. These modules were recently shown to function primarily as beta-mannan-binding modules and have accordingly been classified as members of a novel family of CBMs, family 27. The N-terminal CBM27 (CsCBM27-1) of Man26 from Caldicellulosiruptor Rt8B.4 displays high-binding affinity towards mannohexaose with a Ka of 1 x 10(7) M(-1). Accordingly, the high-resolution crystal structures of CsCBM27-1 native and its mannohexaose complex were solved at 1.55 angstroms and 1.06 angstoms resolution, respectively. In the crystal, CsCBM27-1 shows the typical beta-sandwich jellyroll fold observed in other CBMs with a single metal ion bound, which was identified as calcium. The crystal structures reveal that the overall fold of CsCBM27-1 remains virtually unchanged upon sugar binding and that binding is mediated by three solvent-exposed tryptophan residues and few direct hydrogen bonds. Based on binding affinity and thermal unfolding experiments this structural calcium is shown to play a role in the thermal stability of CsCBM27-1 at high temperatures. The higher binding affinity of CsCBM27-1 to mannooligosaccharides when compared to other members of CBM family 27 might be explained by the different orientation of the residues forming the "aromatic platform" and by differences in the length of loops. Finally, evidence is presented, on the basis of fold similarities and the retention of the position of conserved motifs and a calcium ion, for the consolidation of related CBM families into a superfamily of CBMs.
- Crystallography Group, Max-Delbrück Center for Molecular Medicine, Robert-Rössle-Str. 10, D-13125 Berlin, Germany.
Organizational Affiliation: