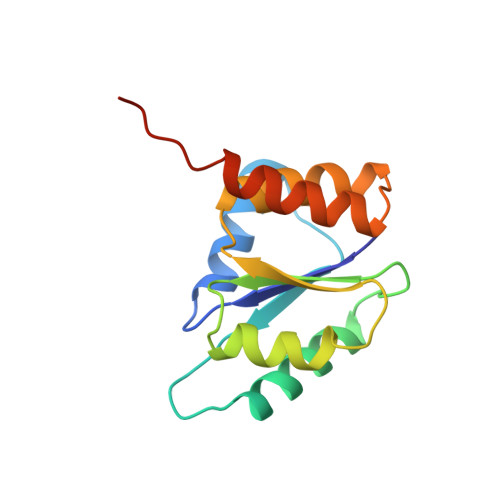
Structure of the IIA domain of the mannose transporter from Escherichia coli at 1.7 angstroms resolution.
Nunn, R.S., Markovic-Housley, Z., Genovesio-Taverne, J.C., Flukiger, K., Rizkallah, P.J., Jansonius, J.N., Schirmer, T., Erni, B.(1996) J Mol Biology 259: 502-511
- PubMed: 8676384 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0335
- Primary Citation Related Structures:
1PDO - PubMed Abstract:
The mannose transporter from Escherichia coli is a member of the phosphoenolpyruvate-dependent phosphotransferase system. The multi-subunit complex couples translocation across the bacterial inner membrane with phosphorylation of the solute. A functional fragment (IIA(Man), residues 2 to 133) of the membrane-associated IIAB(Man) subunit of the mannose transporter was expressed as a selenomethionine protein, and the unphosphorylated molecule was crystallized and its structure solved by X-ray crystallography. The protein consists of a central five-stranded beta-sheet covered by helices on either face. The order of the secondary structure elements is (beta alpha)4, alpha beta. Four beta-strands are arranged in a parallel manner with strand order 2134 and are linked by helices forming right-handed cross-over connections. The fifth strand that forms one edge of the sheet and runs antiparallel to the others is swapped between the subunits of the dimeric structure. Helices D and E form a helical hairpin. Histidine 10, which is transiently phosphorylated during catalysis, is located at the topological switch-point of the structure, close to the subunit interface. Its imidazole ring is hydrogen bonded to the buried side-chain of Asp67. It is likely that Asp67 acts as a general base and thus increases the nucleophilicity of the histidine. Modeling suggests that the covalently bound phosphoryl group would be stabilized by the macrodipole of helix C. Putative interactions between IIA(Man) and the histidine-containing phosphocarrier protein are discussed.
- Biozentrum, Abteilung Strukturbiologie, University of Basel, Switzerland.
Organizational Affiliation: