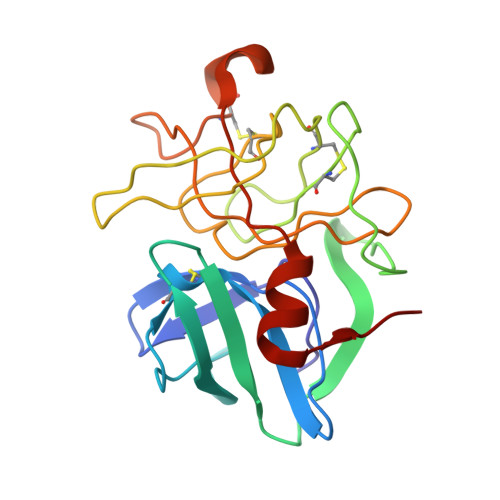
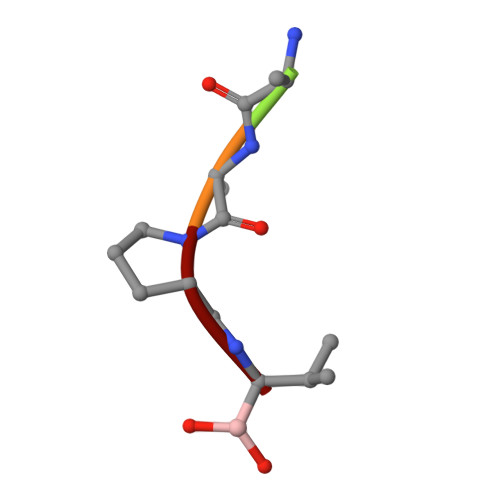
Structural plasticity broadens the specificity of an engineered protease.
Bone, R., Silen, J.L., Agard, D.A.(1989) Nature 339: 191-195
- PubMed: 2716847 Search on PubMed
- DOI: https://doi.org/10.1038/339191a0
- Primary Citation Related Structures:
1P09, 1P10 - PubMed Abstract:
The substrate specificity of alpha-lytic protease has been changed dramatically, with a concomitant increase in activity, by replacing an active-site Met with Ala. The substrate specificity of both this mutant and another similar mutant are extraordinarily broad. X-ray crystallographic analysis shows that structural plasticity, a combination of alternate side-chain conformations and binding-site flexibility, allows both large and small substrates to be well accommodated.
- Department of Biochemistry and Biophysics, University of California, San Francisco 94143-0448.
Organizational Affiliation: