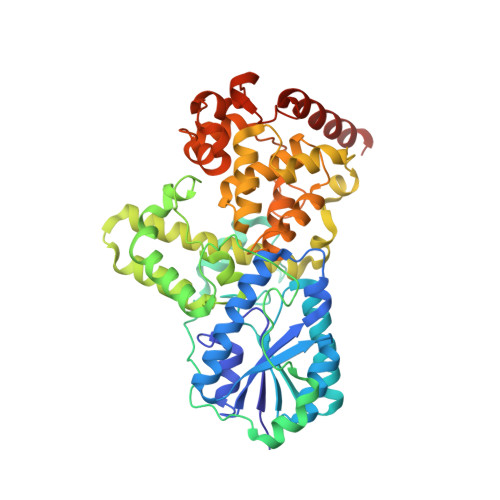
DNA apophotolyase from Anacystis nidulans: 1.8 A structure, 8-HDF reconstitution and X-ray-induced FAD reduction.
Kort, R., Komori, H., Adachi, S., Miki, K., Eker, A.(2004) Acta Crystallogr D Biol Crystallogr 60: 1205-1213
- PubMed: 15213381 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904009321
- Primary Citation Related Structures:
1OWL, 1OWM, 1OWN, 1OWO, 1OWP - PubMed Abstract:
DNA photolyase is a unique flavoenzyme that repairs UV-induced DNA lesions using the energy of visible light. Anacystis nidulans photolyase contains a light-harvesting chromophore, 8-hydroxy-5-deazaflavin (8-HDF), and flavin adenine dinucleotide (FAD) which, in contrast to the 8-HDF chromophore, is indispensable for catalytic activity. This work reports the crystallization and structure at 1.8 A resolution of DNA photolyase devoid of its 8-HDF chromophore (apophotolyase). The overall three-dimensional structure is similar to that of the holoenzyme, indicating that the presence of 8-HDF is not essential for the correct folding of the enzyme. Structural changes include an additional phosphate group, a different conformation for Arg11 and slight rearrangements of Met47, Asp101 and Asp382, which replace part of the 8-HDF molecule in the chromophore-binding pocket. The apophotolyase can be efficiently reconstituted with synthetic 8-hydroxy-5-deazariboflavin, despite the orientation of Arg11 and the presence of the phosphate group in the 8-HDF pocket. Red light or X-rays reduced the FAD chromophore in apophotolyase crystals, as observed by single-crystal spectrophotometry. The structural effects of FAD reduction were determined by comparison of three data sets that were successively collected at 100 K, while the degree of reduction was monitored online by changes in the light absorption of the crystals. X-ray-induced conformational changes were confined to the active site of the protein. They include sub-ångström movements of the O(2) and N(5) atoms of the flavin group as well as the O(delta) atoms of the surrounding amino acids Asp380 and Asn386.
- Laboratory for Microbiology, Swammerdam Institute for Life Sciences, University of Amsterdam, Nieuwe Achtergracht 166, 1018 WV Amsterdam, The Netherlands. rkort@science.uva.nl
Organizational Affiliation: