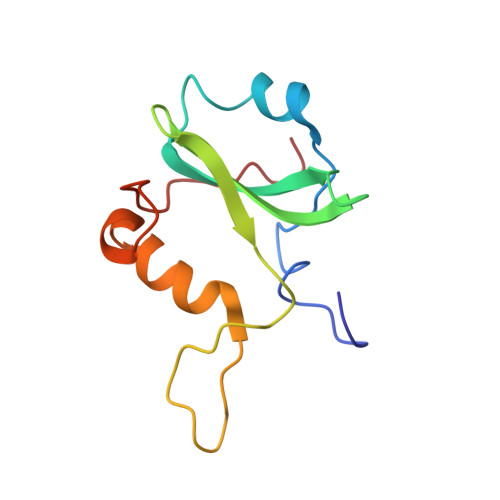
Nuclear magnetic resonance structure of the P395S mutant of the N-SH2 domain of the p85 subunit of PI3 kinase: an SH2 domain with altered specificity
Guenther, U.L., Weyrauch, B., Zhang, X., Schaffhausen, B.(2003) Biochemistry 42: 11120-11127
- PubMed: 14503862 Search on PubMed
- DOI: https://doi.org/10.1021/bi034353x
- Primary Citation Related Structures:
1OO3, 1OO4 - PubMed Abstract:
Understanding the specificity of Src homology 2 (SH2) domains is important because of their critical role in cell signaling. Previous genetic analysis has characterized mutants of the N-terminal src homology 2 (SH2) domain of the p85 subunit of phosphoinositide 3-kinase (PI3K). The P395S mutant exhibits a specificity for phosphopeptide binding different from that of the wild-type SH2. The P395S mutant has an increased affinity for the platelet-derived growth factor receptor (PDGFr) compared to polyomavirus middle T antigen (MT). Solution structures of the P395S mutant of the p85 N-SH2 alone and complexed to a PDGFr phosphopeptide were determined to explain the change in specificity. Chemical shift perturbations caused by different peptides were compared for mutant and wild-type structures. The results show that the single P395S mutation has broad effects on the structure. Furthermore, they provide a rationale for the observed changes in binding preference.
- Institute for Biophysical Chemistry, Centre of Biomolecular Magnetic Resonance, J. W. Goethe University, Frankfurt, Marie-Curie-Strasse 9, 60439 Frankfurt, Germany.
Organizational Affiliation: