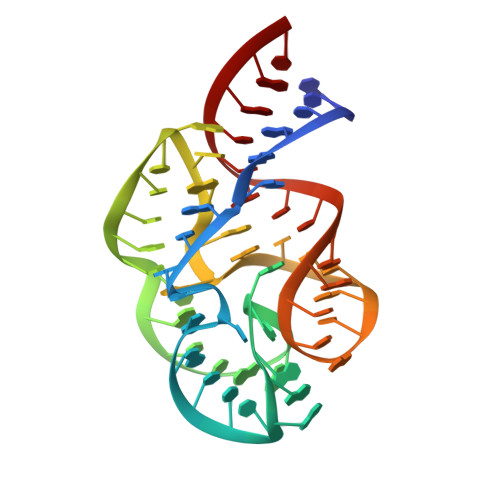
Structural Basis for Contrasting Activities of Ribosome Binding Thiazole Antibiotics
Lentzen, G., Klinck, R., Matassova, N., Aboul-Ela, F., Murchie, A.I.H.(2003) Chem Biol 10: 769
- PubMed: 12954336 Search on PubMed
- DOI: https://doi.org/10.1016/s1074-5521(03)00173-x
- Primary Citation Related Structures:
1OLN - PubMed Abstract:
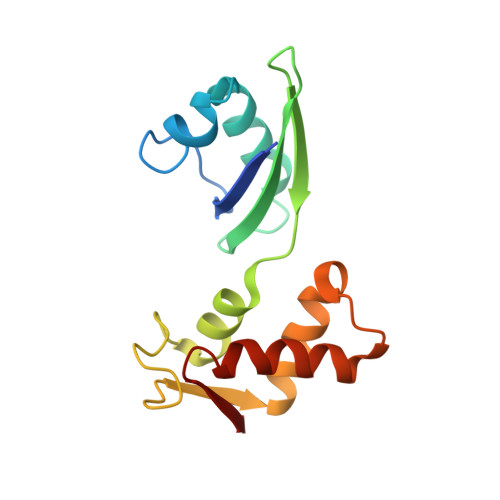
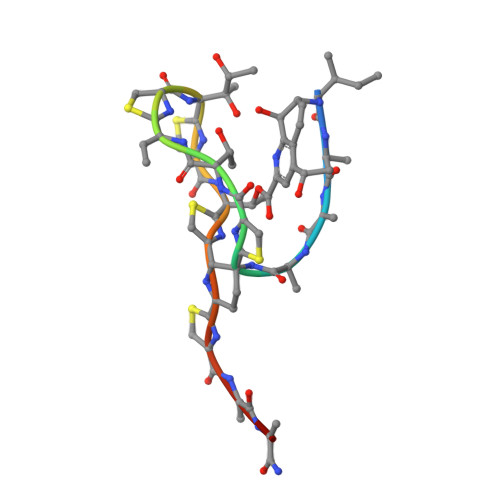
Thiostrepton and micrococcin inhibit protein synthesis by binding to the L11 binding domain (L11BD) of 23S ribosomal RNA. The two compounds are structurally related, yet they produce different effects on ribosomal RNA in footprinting experiments and on elongation factor-G (EF-G)-dependent GTP hydrolysis. Using NMR and an assay based on A1067 methylation by thiostrepton-resistance methyltransferase, we show that the related thiazoles, nosiheptide and siomycin, also bind to this region. The effect of all four antibiotics on EF-G-dependent GTP hydrolysis and EF-G-GDP-ribosome complex formation was studied. Our NMR and biochemical data demonstrate that thiostrepton, nosiheptide, and siomycin share a common profile, which differs from that of micrococcin. We have generated a three-dimensional (3D) model for the interaction of thiostrepton with L11BD RNA. The model rationalizes the differences between micrococcin and the thiostrepton-like antibiotics interacting with L11BD.
- RiboTargets, Ltd., Granta Park, Abington, CB1 6GB, Cambridge, United Kingdom.
Organizational Affiliation: