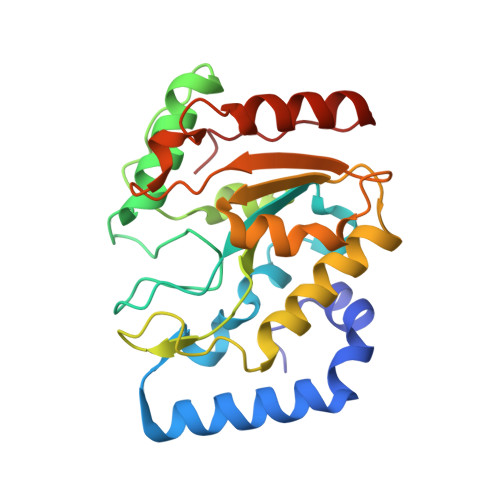
The Crystal Structure of Uracil-DNA Glycosylase from Atlantic Cod (Gadus Morhua) Reveals Cold-Adaptation Features
Leiros, I., Moe, E., Lanes, O., Smalas, A.O., Willassen, N.P.(2003) Acta Crystallogr D Biol Crystallogr 59: 1357
- PubMed: 12876336
- DOI: https://doi.org/10.1107/s0907444903011144
- Primary Citation Related Structures:
1OKB - PubMed Abstract:
Uracil-DNA glycosylase (UDG; EC 3.2.2.3) is a DNA-repair protein that catalyses the hydrolysis of promutagenic uracil residues from single- or double-stranded DNA, generating free uracil and abasic DNA. The crystal structure of the catalytic domain of cod uracil-DNA glycosylase (cUDG) has been determined to 1.9 A resolution, with final R factors of 18.61 and 20.57% for the working and test sets of reflections, respectively. This is the first crystal structure of a uracil-DNA glycosylase from a cold-adapted species and a detailed comparison with the human enzyme is performed in order to rationalize the cold-adapted behaviour of the cod enzyme at the structural level. The catalytic domain of cUDG comprises 223 residues, with a sequence identity to the human UDG of 75%. The tertiary structures of the two enzymes are also similar, with an overall displacement in main-chain atomic positions of 0.63 A. The amino-acid substitutions and the differences in intramolecular hydrogen bonds, hydrophobic interactions, ion-pair interactions and electrostatic potentials are compared and discussed in order to gain insight into the factors that cause the increased activity and reduced thermostability of the cod enzyme. In particular, the reduced number of strong ion-pair interactions in the C-terminal half of cUDG is believed to greatly affect the flexibility and/or stability. Increased positive electrostatic surface potential on the DNA-facing side of cUDG seems to be responsible for increasing the affinity for the negatively charged DNA compared with that of hUDG.
- Department of Chemistry, Faculty of Science, University of Tromsø, N-9037 Tromsø, Norway.
Organizational Affiliation: