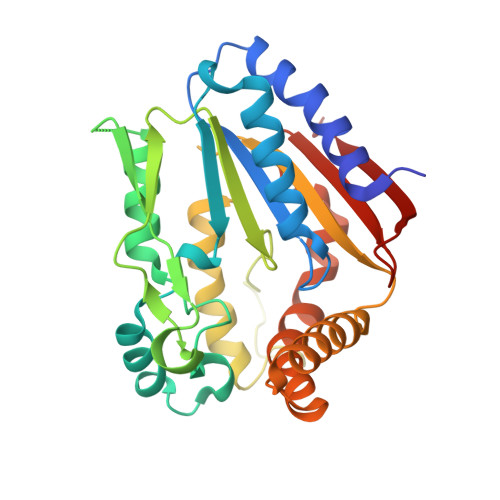
Crystal Structure of the Ygr205W Protein from Saccharomyces Cerevisiae: Close Structural Resemblance to E.Coli Pantothenate Kinase
Li De La Sierra-Gallay, I., Collinet, B., Graille, M., Quevillon-Cheruel, S., Liger, D., Minard, P., Blondeau, K., Henckes, G., Aufrere, R., Leulliot, N., Zhou, C.Z., Sorrel, I., Ferrer, J.L., Poupon, A., Janin, J., Van Tilbeurgh, H.(2004) Proteins 54: 776
- PubMed: 14997573
- DOI: https://doi.org/10.1002/prot.10596
- Primary Citation Related Structures:
1ODF - PubMed Abstract:
The protein product of the YGR205w gene of Saccharomyces cerevisiae was targeted as part of our yeast structural genomics project. YGR205w codes for a small (290 amino acids) protein with unknown structure and function. The only recognizable sequence feature is the presence of a Walker A motif (P loop) indicating a possible nucleotide binding/converting function. We determined the three-dimensional crystal structure of Se-methionine substituted protein using multiple anomalous diffraction. The structure revealed a well known mononucleotide fold and strong resemblance to the structure of small metabolite phosphorylating enzymes such as pantothenate and phosphoribulo kinase. Biochemical experiments show that YGR205w binds specifically ATP and, less tightly, ADP. The structure also revealed the presence of two bound sulphate ions, occupying opposite niches in a canyon that corresponds to the active site of the protein. One sulphate is bound to the P-loop in a position that corresponds to the position of beta-phosphate in mononucleotide protein ATP complex, suggesting the protein is indeed a kinase. The nature of the phosphate accepting substrate remains to be determined.
- Laboratoire d'Enzymologie et Biochimie Structurales (CNRS-UPR 9063), Bât. 34, 1 Av. de la Terrasse, 91198 Gif sur Yvette, France.
Organizational Affiliation: