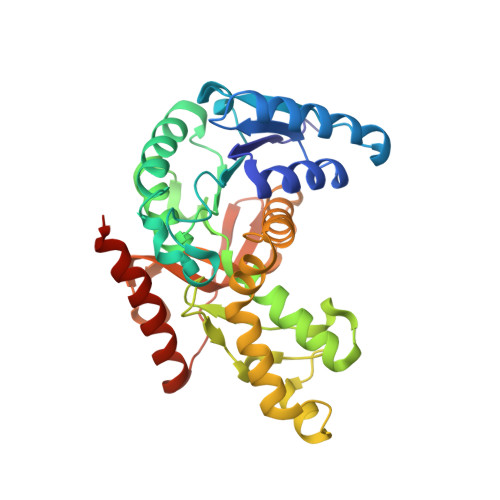
Crystal Structure of Plasmodium Berghei Lactate Dehydrogenase Indicates the Unique Structural Differences of These Enzymes are Shared Across the Plasmodium Genus
Winter, V.J., Cameron, A., Tranter, R., Sessions, R.B., Brady, R.L.(2003) Mol Biochem Parasitol 131: 1
- PubMed: 12967707 Search on PubMed
- DOI: https://doi.org/10.1016/s0166-6851(03)00170-1
- Primary Citation Related Structures:
1OC4 - PubMed Abstract:
As Plasmodium rely extensively on homolactic fermentation for energy production, Plasmodium falciparum lactate dehydrogenase (PfLDH)--the key enzyme in this process--has previously been suggested as a novel target for antimalarials. This enzyme has distinctive kinetic and structural properties that distinguish it from its human homologues. In this study, we now describe the expression, kinetic characterisation and crystal structure determination of the LDH from Plasmodium berghei. This enzyme is seen to have a similar kinetic profile to its P. falciparum counterpart, exhibiting the characteristic lack of substrate inhibition that distinguishes plasmodial from human LDHs. The crystal structure of P. berghei lactate dehydrogenase (PbLDH) shows a very similar active site arrangement to the P. falciparum enzyme. In particular, an insertion of five amino acid residues in the active site loop creates an enlarged volume in the substrate binding site, and characteristic changes in the residues lining the NADH cofactor binding pocket result in displacement of the cofactor relative to its observed position in mammalian and all other LDH structures. These results imply the special features previously described for PfLDH may be shared across the Plasmodium genus, supporting the universal application of therapeutics targeting this enzyme.
- Department of Biochemistry, University of Bristol, Bristol BS8 1TD, UK.
Organizational Affiliation: