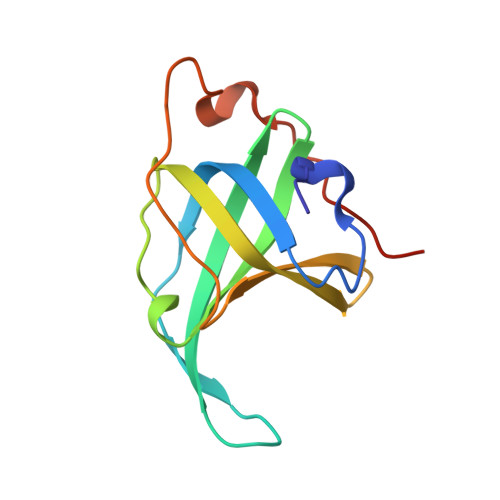
Insights into ssDNA recognition by the OB fold from a structural and thermodynamic study of Sulfolobus SSB protein.
Kerr, I.D., Wadsworth, R.I., Cubeddu, L., Blankenfeldt, W., Naismith, J.H., White, M.F.(2003) EMBO J 22: 2561-2570
- PubMed: 12773373 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/cdg272
- Primary Citation Related Structures:
1O7I - PubMed Abstract:
Information processing pathways such as DNA replication are conserved in eukaryotes and archaea and are significantly different from those found in bacteria. Single-stranded DNA-binding (SSB) proteins (or replication protein A, RPA, in eukaryotes) play a central role in many of these pathways. However, whilst euryarchaea have a eukaryotic-type RPA homologue, crenarchaeal SSB proteins appear much more similar to the bacterial proteins, with a single OB fold for DNA binding and a flexible C-terminal tail that is implicated in protein-protein interactions. We have determined the crystal structure of the SSB protein from the crenarchaeote Sulfolobus solfataricus to 1.26 A. The structure shows a striking and unexpected similarity to the DNA-binding domains of human RPA, providing confirmation of the close relationship between archaea and eukaryotes. The high resolution of the structure, together with thermodynamic and mutational studies of DNA binding, allow us to propose a molecular basis for DNA binding and define the features required for eukaryotic and archaeal OB folds.
- Centre for Biomolecular Science, St Andrews University, Fife, KY16 9ST, UK.
Organizational Affiliation: