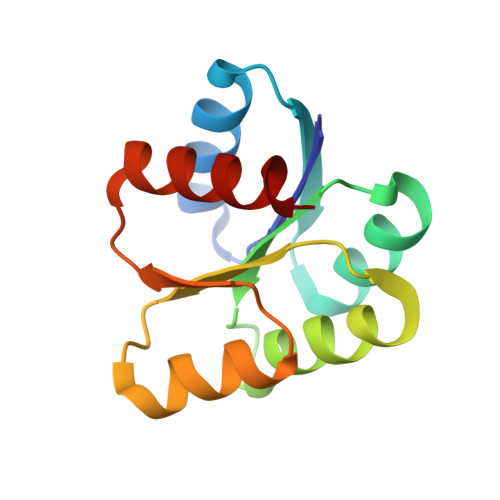
Crystal structure of the response regulator 02 receiver domain, the essential YycF two-component system of Streptococcus pneumoniae in both complexed and native states.
Bent, C.J., Isaacs, N.W., Mitchell, T.J., Riboldi-Tunnicliffe, A.(2004) J Bacteriol 186: 2872-2879
- PubMed: 15090529 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.186.9.2872-2879.2004
- Primary Citation Related Structures:
1NXO, 1NXP, 1NXT, 1NXW - PubMed Abstract:
A variety of bacterial cellular responses to environmental signals are mediated by two-component signal transduction systems comprising a membrane-associated histidine protein kinase and a cytoplasmic response regulator (RR), which interpret specific stimuli and produce a measured physiological response. In RR activation, transient phosphorylation of a highly conserved aspartic acid residue drives the conformation changes needed for full activation of the protein. Sequence homology reveals that RR02 from Streptococcus pneumoniae belongs to the OmpR subfamily of RRs. The structures of the receiver domains from four members of this family, DrrB and DrrD from Thermotoga maritima, PhoB from Escherichia coli, and PhoP from Bacillus subtilis, have been elucidated. These domains are globally very similar in that they are composed of a doubly wound alpha(5)beta(5); however, they differ remarkably in the fine detail of the beta4-alpha4 and alpha4 regions. The structures presented here reveal a further difference of the geometry in this region. RR02 is has been shown to be the essential RR in the gram-positive bacterium S. pneumoniae R. Lange, C. Wagner, A. de Saizieu, N. Flint, J. Molnos, M. Stieger, P. Caspers, M. Kamber, W. Keck, and K. E. Amrein, Gene 237:223-234, 1999; J. P. Throup, K. K. Koretke, A. P. Bryant, K. A. Ingraham, A. F. Chalker, Y. Ge, A. Marra, N. G. Wallis, J. R. Brown, D. J. Holmes, M. Rosenberg, and M. K. Burnham, Mol. Microbiol. 35:566-576, 2000). RR02 functions as part of a phosphotransfer system that ultimately controls the levels of competence within the bacteria. Here we report the native structure of the receiver domain of RR02 from serotype 4 S. pneumoniae (as well as acetate- and phosphate-bound forms) at different pH levels. Two native structures at 2.3 A, phased by single-wavelength anomalous diffraction (xenon SAD), and 1.85 A and a third structure at pH 5.9 revealed the presence of a phosphate ion outside the active site. The fourth structure revealed the presence of an acetate molecule in the active site.
- Department of Chemistry, Division of Infection and Immunity, University of Glasgow, Glasgow G12 8QQ, Scotland, United Kingdom.
Organizational Affiliation: