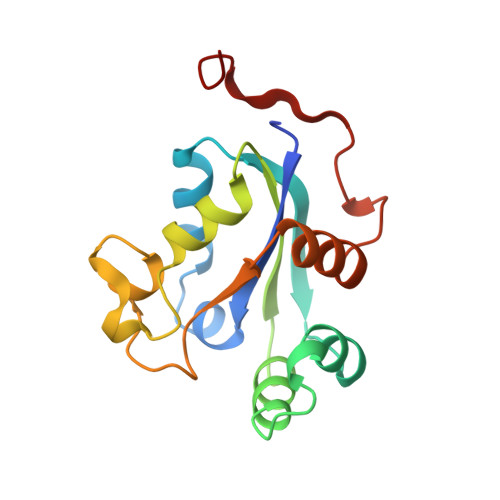
Refined X-ray structure of Dictyostelium discoideum nucleoside diphosphate kinase at 1.8 A resolution.
Morera, S., LeBras, G., Lascu, I., Lacombe, M.L., Veron, M., Janin, J.(1994) J Mol Biology 243: 873-890
- PubMed: 7966307
- DOI: https://doi.org/10.1006/jmbi.1994.1689
- Primary Citation Related Structures:
1NPK - PubMed Abstract:
The X-ray structure of the nucleoside diphosphate kinase (NDP kinase) from Dictyostelium discoideum has been refined at 1.8 A resolution from a hexagonal crystal form with a 17 kDa monomer in its asymmetric unit. The atomic model was derived from the previously determined structure of a point mutant of the protein. It contains 150 amino acid residues out of 155, and 95 solvent molecules. The R-factor is 0.196 and the estimated accuracy of the average atomic position, 0.25 A. The Dictyostelium structure is described in detail and compared to those of Drosophila and Myxococcus xanthus NDP kinases. The protein is a hexamer with D3 symmetry. Residues 8 to 138 of each subunit form a globular alpha/beta domain. The four-stranded beta-sheet is antiparallel; its topology is different from other phosphate transfer enzymes, and also from the HPr protein which, like NDP kinase, carries a phosphorylated histidine. The same topology is nevertheless found in several other proteins that bind mononucleotides, RNA or DNA. Strand connections in NDP kinase involve alpha-helices and a 20-residue segment called the Kpn loop. The beta-sheet is regular except for a beta-bulge in edge strand beta 2 and a gamma-turn at residue Ile120 just preceding strand beta 4. The latter may induce strain in the main chain near the active site His122. The alpha 1 beta 2 motif participates in forming dimers within the hexamer, helices alpha 1 and alpha 3, the Kpn loop and C terminus, in forming trimers. The subunit fold and dimer interactions found in Dictyostelium are conserved in other NDP kinases. Trimer interactions probably occur in all eukaryotic enzymes. They are absent in the bacterial Myxococcus xanthus enzyme which is a tetramer, even though the subunit structure is very similar. In Dictyostelium, contacts between Kpn loops near the 3-fold axis block access to a central cavity lined with polar residues and filled with well-defined solvent molecules. Biochemical data on point mutants highlight the contribution of the Kpn loop to protein stability. In Myxococcus, the Kpn loops are on the tetramer surface and their sequence is poorly conserved. Yet, their conformation is maintained and they make a similar contribution to the substrate binding site.
- Laboratoire de Biologie Structurale, UMR 9920 CNRS-Université Paris-Sud, Gif-sur-Yvette, France.
Organizational Affiliation: