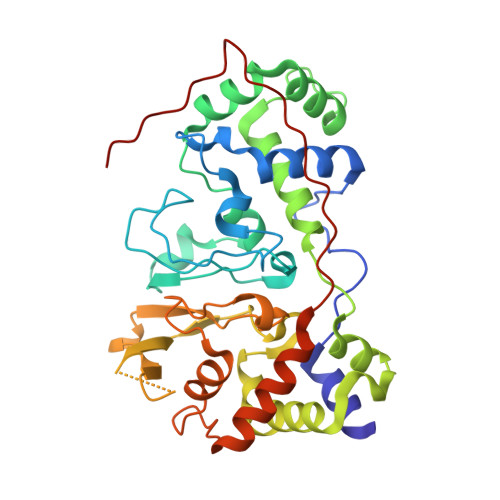
Structural basis for the mechanism of Ca(2+) activation of the di-heme cytochrome c peroxidase from Pseudomonas nautica 617
Dias, J.M., Alves, T., Pereira, A.S., Bourgeois, D., Moura, I.(2004) Structure 12: 961-973
- PubMed: 15274917 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.03.025
- Primary Citation Related Structures:
1NML, 1RZ5, 1RZ6 - PubMed Abstract:
Cytochrome c peroxidase (CCP) catalyses the reduction of H(2)O(2) to H(2)O, an important step in the cellular detoxification process. The crystal structure of the di-heme CCP from Pseudomonas nautica 617 was obtained in two different conformations in a redox state with the electron transfer heme reduced. Form IN, obtained at pH 4.0, does not contain Ca(2+) and was refined at 2.2 A resolution. This inactive form presents a closed conformation where the peroxidatic heme adopts a six-ligand coordination, hindering the peroxidatic reaction from taking place. Form OUT is Ca(2+) dependent and was crystallized at pH 5.3 and refined at 2.4 A resolution. This active form shows an open conformation, with release of the distal histidine (His71) ligand, providing peroxide access to the active site. This is the first time that the active and inactive states are reported for a di-heme peroxidase.
- REQUIMTE/CQFB, Departamento de Química, FCT, Universidade Nova de Lisboa, 2829-516 Caparica, Portugal.
Organizational Affiliation: