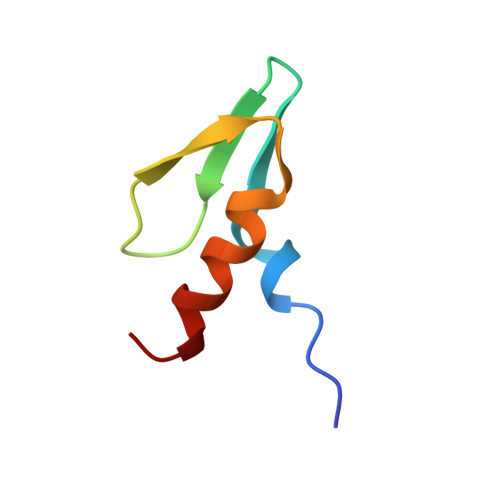
The solution structure of the first zinc finger domain of SWI5: a novel structural extension to a common fold.
Dutnall, R.N., Neuhaus, D., Rhodes, D.(1996) Structure 4: 599-611
- PubMed: 8736557
- DOI: https://doi.org/10.1016/s0969-2126(96)00064-0
- Primary Citation of Related Structures:
1NCS - PubMed Abstract:
The 2Cys-2His (C2-H2) zinc finger is a protein domain commonly used for sequence-specific DNA recognition. The zinc fingers of the yeast transcription factors SWI5 and ACE2 share strong sequence homology, which extends into a region N-terminal to the first finger, suggesting that the DNA-binding domains of these two proteins include additional structural elements. Structural analysis of the zinc fingers of SWI5 reveals that a 15 residue region N-terminal to the finger motifs forms part of the structure of the first finger domain, adding a beta strand and a helix not previously observed in other zinc finger structures. Sequence analysis suggests that other zinc finger proteins may also have this structure. Biochemical studies show that this additional structure increases DNA-binding affinity. The structural analysis presented reveals a novel zinc finger structure in which additional structural elements have been added to the C2-H2 zinc finger fold. This additional structure may enhance stability and has implications for DNA recognition by extending the potential DNA-binding surface of a single zinc finger domain.
- MRC Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: