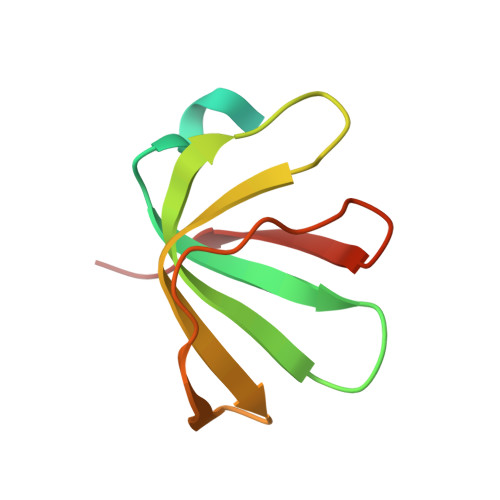
Homomeric ring assemblies of eukaryotic Sm proteins have affinity for both RNA and DNA: Crystal structure of an oligomeric complex of yeast SmF
Collins, B.M., Cubeddu, L., Naidoo, N., Harrop, S.J., Kornfeld, G.D., Dawes, I.W., Curmi, P.M.G., Mabbutt, B.C.(2003) J Biological Chem 278: 17291-17298
- PubMed: 12618433
- DOI: https://doi.org/10.1074/jbc.M211826200
- Primary Citation Related Structures:
1N9R, 1N9S - PubMed Abstract:
Sm and Sm-like proteins are key components of small ribonucleoproteins involved in many RNA and DNA processing pathways. In eukaryotes, these complexes contain seven unique Sm or Sm-like (Lsm) proteins assembled as hetero-heptameric rings, whereas in Archaea and bacteria six or seven-membered rings are made from only a single polypeptide chain. Here we show that single Sm and Lsm proteins from yeast also have the capacity to assemble into homo-oligomeric rings. Formation of homo-oligomers by the spliceosomal small nuclear ribonucleoprotein components SmE and SmF preclude hetero-interactions vital to formation of functional small nuclear RNP complexes in vivo. To better understand these unusual complexes, we have determined the crystal structure of the homomeric assembly of the spliceosomal protein SmF. Like its archaeal/bacterial homologs, the SmF complex forms a homomeric ring but in an entirely novel arrangement whereby two heptameric rings form a co-axially stacked dimer via interactions mediated by the variable loops of the individual SmF protein chains. Furthermore, we demonstrate that the homomeric assemblies of yeast Sm and Lsm proteins are capable of binding not only to oligo(U) RNA but, in the case of SmF, also to oligo(dT) single-stranded DNA.
- Cambridge Institute for Medical Research, University of Cambridge, Department of Clinical Biochemistry, Hills Road, Cambridge CB2 2XY, United Kingdom.
Organizational Affiliation: