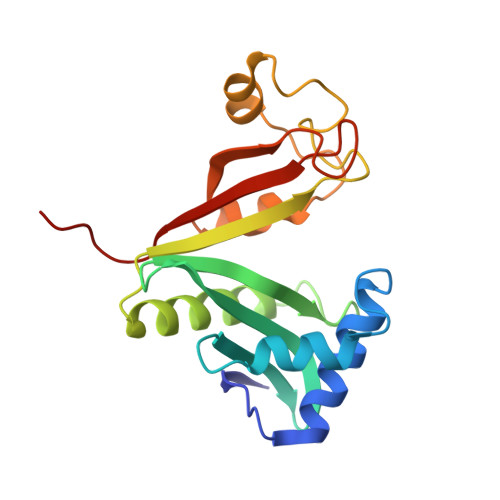
X-ray structure of the AAC(6')-Ii antibiotic resistance enzyme at 1.8 A resolution; examination of oligomeric arrangements in GNAT superfamily members
Burk, D.L., Ghuman, N., Wybenga-Groot, L.E., Berghuis, A.M.(2003) Protein Sci 12: 426-437
- PubMed: 12592013 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.0233503
- Primary Citation Related Structures:
1N71 - PubMed Abstract:
The rise of antibiotic resistance as a public health concern has led to increased interest in studying the ways in which bacteria avoid the effects of antibiotics. Enzymatic inactivation by several families of enzymes has been observed to be the predominant mechanism of resistance to aminoglycoside antibiotics such as kanamycin and gentamicin. Despite the importance of acetyltransferases in bacterial resistance to aminoglycoside antibiotics, relatively little is known about their structure and mechanism. Here we report the three-dimensional atomic structure of the aminoglycoside acetyltransferase AAC(6')-Ii in complex with coenzyme A (CoA). This structure unambiguously identifies the physiologically relevant AAC(6')-Ii dimer species, and reveals that the enzyme structure is similar in the AcCoA and CoA bound forms. AAC(6')-Ii is a member of the GCN5-related N-acetyltransferase (GNAT) superfamily of acetyltransferases, a diverse group of enzymes that possess a conserved structural motif, despite low sequence homology. AAC(6')-Ii is also a member of a subset of enzymes in the GNAT superfamily that form multimeric complexes. The dimer arrangements within the multimeric GNAT superfamily members are compared, revealing that AAC(6')-Ii forms a dimer assembly that is different from that observed in the other multimeric GNAT superfamily members. This different assembly may provide insight into the evolutionary processes governing dimer formation.
- Departments of Biochemistry and Microbiology and Immunology, McGill University, Montreal, Quebec H3A 2B4, Canada.
Organizational Affiliation: