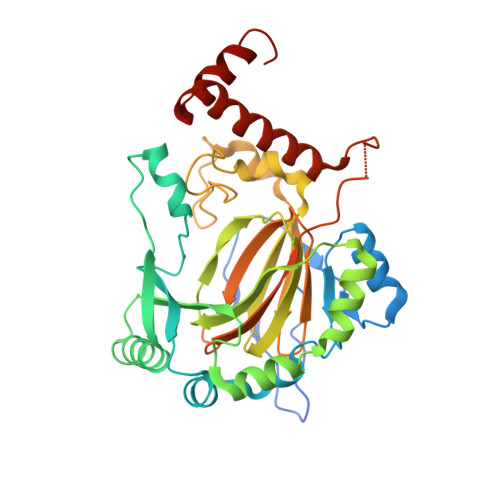
Structure of Factor-Inhibiting Hypoxia-Inducible Factor 1: An Asparaginyl Hydroxylase Involved in the Hypoxic Response Pathway
Dann III, C.E., Bruick, R.K., Deisenhofer, J.(2002) Proc Natl Acad Sci U S A 99: 15351-15356
- PubMed: 12432100
- DOI: https://doi.org/10.1073/pnas.202614999
- Primary Citation of Related Structures:
1MZE, 1MZF - PubMed Abstract:
Precise regulation of the evolutionarily conserved hypoxia-inducible transcription factor (HIF) ensures proper adaptation to variations in oxygen availability throughout development and into adulthood. Oxygen-dependent regulation of HIF stability and activity are mediated by hydroxylation of conserved proline and asparagine residues, respectively. Because the relevant prolyl and asparginyl hydroxylases use O(2) to effect these posttranslational modifications, these enzymes are implicated as direct oxygen sensors in the mammalian hypoxic response pathway. Here we present the structure of factor-inhibiting HIF-1 (FIH-1), the pertinent asparaginyl hydroxylase involved in hypoxic signaling. Hydroxylation of the C-terminal transactivation domain (CTAD) of HIF by FIH-1 prevents CTAD association with transcriptional coactivators under normoxic conditions. Consistent with other structurally known hydroxylases, FIH-1 is comprised of a beta-strand jellyroll core with both Fe(II) and the cosubstrate 2-oxoglutarate bound in the active site. Details of the molecular contacts at the active site of FIH-1 have been elucidated and provide a platform for future drug design. Furthermore, the structure reveals the presence of a FIH-1 homodimer that forms in solution and is essential for FIH activity.
- Howard Hughes Medical Institute and Department of Biochemistry, University of Texas Southwestern Medical Center, 5323 Harry Hines Boulevard, Dallas 75390, USA.
Organizational Affiliation: