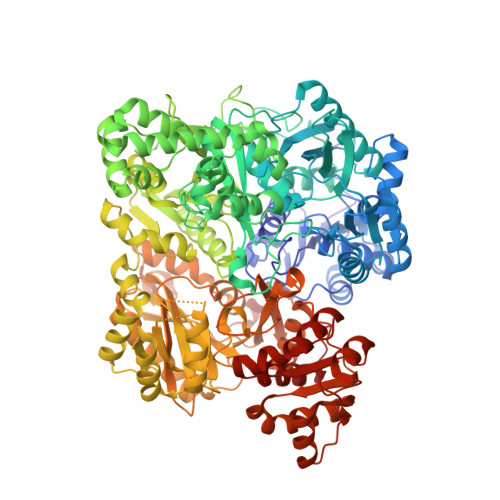
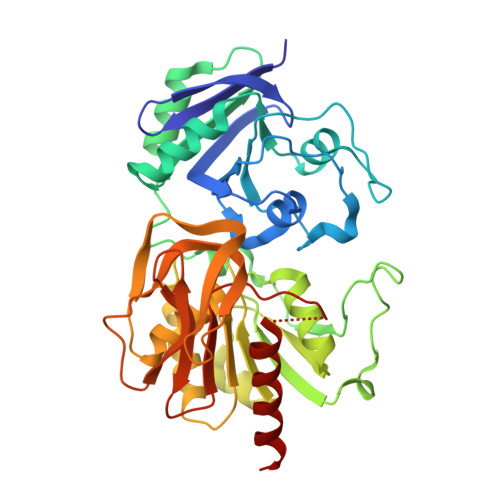
Carbamoyl-phosphate synthetase. Creation of an escape route for ammonia
Thoden, J.B., Huang, X., Raushel, F.M., Holden, H.M.(2002) J Biological Chem 277: 39722-39727
- PubMed: 12130656 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M206915200
- Primary Citation Related Structures:
1M6V - PubMed Abstract:
Carbamoyl-phosphate synthetase catalyzes the production of carbamoyl phosphate through a reaction mechanism requiring one molecule of bicarbonate, two molecules of MgATP, and one molecule of glutamine. The enzyme from Escherichia coli is composed of two polypeptide chains. The smaller of these belongs to the Class I amidotransferase superfamily and contains all of the necessary amino acid side chains required for the hydrolysis of glutamine to glutamate and ammonia. Two homologous domains from the larger subunit adopt conformations that are characteristic for members of the ATP-grasp superfamily. Each of these ATP-grasp domains contains an active site responsible for binding one molecule of MgATP. High resolution x-ray crystallographic analyses have shown that, remarkably, the three active sites in the E. coli enzyme are connected by a molecular tunnel of approximately 100 A in total length. Here we describe the high resolution x-ray crystallographic structure of the G359F (small subunit) mutant protein of carbamoyl phosphate synthetase. This residue was initially targeted for study because it resides within the interior wall of the molecular tunnel leading from the active site of the small subunit to the first active site of the large subunit. It was anticipated that a mutation to the larger residue would "clog" the ammonia tunnel and impede the delivery of ammonia from its site of production to the site of utilization. In fact, the G359F substitution resulted in a complete change in the conformation of the loop delineated by Glu-355 to Ala-364, thereby providing an "escape" route for the ammonia intermediate directly to the bulk solvent. The substitution also effected the disposition of several key catalytic amino acid side chains in the small subunit active site.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin, 53706-1544, USA.
Organizational Affiliation: