Residues involved in the catalysis, base specificity, and cytotoxicity of ribonuclease from Rana catesbeiana based upon mutagenesis and X-ray crystallography
Leu, Y.-J., Chern, S.-S., Wang, S.-C., Hsiao, Y.-Y., Amiraslanov, I., Liaw, Y.-C., Liao, Y.-D.(2003) J Biological Chem 278: 7300-7309
- PubMed: 12499382 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M206701200
- Primary Citation Related Structures:
1M07 - PubMed Abstract:
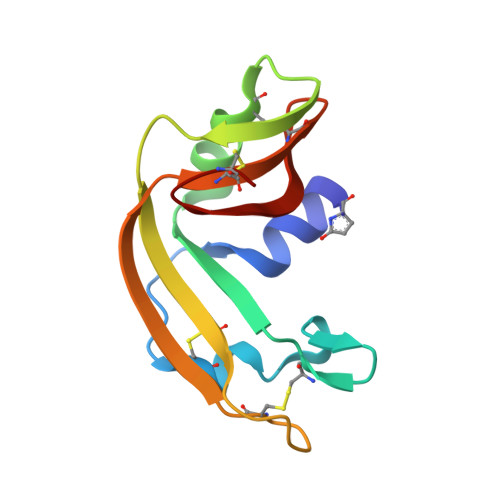
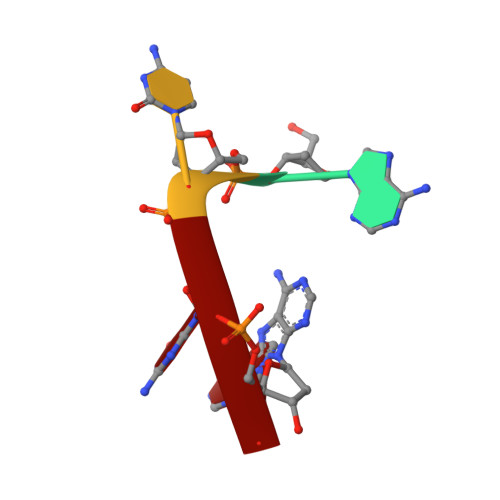
The Rana catesbeiana (bullfrog) ribonucleases, which belong to the RNase A superfamily, exert cytotoxicity toward tumor cells. RC-RNase, the most active among frog ribonucleases, has a unique base preference for pyrimidine-guanine rather than pyrimidine-adenine in RNase A. Residues of RC-RNase involved in base specificity and catalytic activity were determined by site-directed mutagenesis, k(cat)/K(m) analysis toward dinucleotides, and cleavage site analysis of RNA substrate. The results show that Pyr-1 (N-terminal pyroglutamate), Lys-9, and Asn-38 along with His-10, Lys-35, and His-103 are involved in catalytic activity, whereas Pyr-1, Thr-39, Thr-70, Lys-95, and Glu-97 are involved in base specificity. The cytotoxicity of RC-RNase is correlated, but not proportional to, its catalytic activity. The crystal structure of the RC-RNase.d(ACGA) complex was determined at 1.80 A resolution. Residues Lys-9, His-10, Lys-35, and His-103 interacted directly with catalytic phosphate at the P(1) site, and Lys-9 was stabilized by hydrogen bonds contributed by Pyr-1, Tyr-28, and Asn-38. Thr-70 acts as a hydrogen bond donor for cytosine through Thr-39 and determines B(1) base specificity. Interestingly, Pyr-1 along with Lys-95 and Glu-97 form four hydrogen bonds with guanine at B(2) site and determine B(2) base specificity.
- Institute of Biomedical Sciences and the Institute of Molecular Biology, Academia Sinica, Taipei 115, and the Department of Life Science, National Tsing-Hua University, Hsin-Chu 300, Taiwan.
Organizational Affiliation: